Development of a protein marker panel for characterization of human induced pluripotent stem cells (hiPSCs) using global quantitative proteome analysis
- PMID: 25840413
- PMCID: PMC5778352
- DOI: 10.1016/j.scr.2015.01.009
Development of a protein marker panel for characterization of human induced pluripotent stem cells (hiPSCs) using global quantitative proteome analysis
Abstract
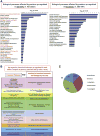
The emergence of new methods for reprogramming of adult somatic cells into induced pluripotent stem cells (iPSC) led to the development of new approaches in drug discovery and regenerative medicine. Investigation of the molecular mechanisms underlying the self-renewal, expansion and differentiation of human iPSC (hiPSC) should lead to improvements in the manufacture of safe and reliable cell therapy products. The goal of our study was qualitative and quantitative proteomic characterizations of hiPSC by means of electrospray ionization (ESI)-MS(e) and MALDI-TOF/TOF mass spectrometry (MS). Proteomes of hiPSCs of different somatic origins: fibroblasts and peripheral blood CD34(+) cells, reprogrammed by the same technique, were compared with the original somatic cells and hESC. Quantitative proteomic comparison revealed approximately 220 proteins commonly up-regulated in all three pluripotent stem cell lines compared to the primary cells. Expression of 21 proteins previously reported as pluripotency markers was up-regulated in both hiPSCs (8 were confirmed by Western blot). A number of novel candidate marker proteins with the highest fold-change difference between hiPSCs/hESC and somatic cells discovered by MS were confirmed by Western blot. A panel of 22 candidate marker proteins of hiPSC was developed and expression of these proteins was confirmed in 8 additional hiPSC lines.
Published by Elsevier B.V.
Conflict of interest statement
There is no conflict of interest for any author to report.
Figures




References
-
- Bachy I, Franck MC, Li L, Abdo H, Pattyn A, Ernfors P. The transcription factor Cux2 marks development of an A-delta sublineage of TrkA sensory neurons. Dev. Biol. 2011;360:77–86. - PubMed
-
- Bar-Nur O, Russ HA, Efrat S, Benvenisty N. Epigenetic memory and preferential lineage-specific differentiation in induced pluripotent stem cells derived from human pancreatic islet beta cells. Cell Stem Cell. 2011;9:17–23. - PubMed
-
- Benevento M, Munoz J. Role of mass spectrometry-based proteomics in the study of cellular reprogramming and induced pluripotent stem cells. Expert Rev. Proteomics. 2012;9:379–399. - PubMed
-
- Bodnar WM, Blackburn RK, Krise JM, Moseley MA. Exploiting the complementary nature of LC/MALDI/MS/MS and LC/ESI/MS/MS for increased proteome coverage. J. Am. Soc. Mass Spectrom. 2003;14(9):971–979. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

