A pyrosequencing insight into sprawling bacterial diversity and community dynamics in decaying deadwood logs of Fagus sylvatica and Picea abies
- PMID: 25851097
- PMCID: PMC4389208
- DOI: 10.1038/srep09456
A pyrosequencing insight into sprawling bacterial diversity and community dynamics in decaying deadwood logs of Fagus sylvatica and Picea abies
Erratum in
-
Erratum: A pyrosequencing insight into sprawling bacterial diversity and community dynamics in decaying deadwood logs of Fagus sylvatica and Picea abies.Sci Rep. 2016 May 10;6:10498. doi: 10.1038/srep10498. Sci Rep. 2016. PMID: 27162103 Free PMC article. No abstract available.
Abstract
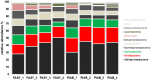
Deadwood is an important biodiversity hotspot in forest ecosystems. While saproxylic insects and wood-inhabiting fungi have been studied extensively, little is known about deadwood-inhabiting bacteria. The study we present is among the first to compare bacterial diversity and community structure of deadwood under field conditions. We therefore compared deadwood logs of two temperate forest tree species Fagus sylvatica and Picea abies using 16S rDNA pyrosequencing to identify changes in bacterial diversity and community structure at different stages of decay in forest plots under different management regimes. Alphaproteobacteria, Acidobacteria and Actinobacteria were the dominant taxonomic groups in both tree species. There were no differences in bacterial OTU richness between deadwood of Fagus sylvatica and Picea abies. Bacteria from the order Rhizobiales became more abundant during the intermediate and advanced stages of decay, accounting for up to 25% of the entire bacterial community in such logs. The most dominant OTU was taxonomically assigned to the genus Methylovirgula, which was recently described in a woodblock experiment of Fagus sylvatica. Besides tree species we were able to demonstrate that deadwood physico-chemical properties, in particular remaining mass, relative wood moisture, pH, and C/N ratio serve as drivers of community composition of deadwood-inhabiting bacteria.
Figures


References
-
- Harmon M. E. et al. Ecology of coarse woody debris in temperate ecosystems. Adv. Ecol. Res. 15, 133–302 (1986).
-
- Lassauce A., Paillet Y., Jactel H. & Bouget C. Deadwood as a surrogate for forest biodiversity: Meta-analysis of correlations between deadwood volume and species richness of saproxylic organisms. Ecol. Indic. 11, 1027–1039 (2011).
-
- Stokland J. N., Siitonen J. & Jonsson B. G. Biodiversity in Dead Wood. [1–509] (Cambridge University Press, Cambridge, 2012).
-
- Cornwell W. K. et al. Plant traits and wood fates across the globe: rotted, burned, or consumed? Glob. Change Biol. 15, 2431–2449 (2009).
-
- Kahl T., Mund M., Bauhus J. & Schulze E. D. Dissolved organic carbon from European beech logs: Patterns of input to and retention by surface soil. Ecoscience 19, 1–10 (2012).
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources

