GRP78/Dna K Is a Target for Nexavar/Stivarga/Votrient in the Treatment of Human Malignancies, Viral Infections and Bacterial Diseases
- PMID: 25858032
- PMCID: PMC4843173
- DOI: 10.1002/jcp.25014
GRP78/Dna K Is a Target for Nexavar/Stivarga/Votrient in the Treatment of Human Malignancies, Viral Infections and Bacterial Diseases
Abstract
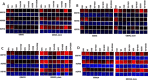
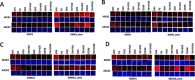
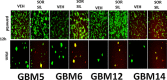
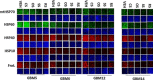
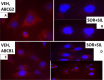
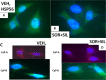
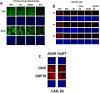
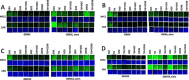
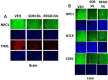
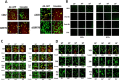
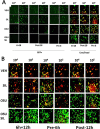
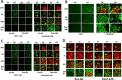
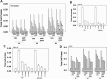
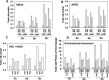
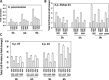
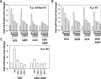
Prior tumor cell studies have shown that the drugs sorafenib (Nexavar) and regorafenib (Stivarga) reduce expression of the chaperone GRP78. Sorafenib/regorafenib and the multi-kinase inhibitor pazopanib (Votrient) interacted with sildenafil (Viagra) to further rapidly reduce GRP78 levels in eukaryotes and as single agents to reduce Dna K levels in prokaryotes. Similar data were obtained in tumor cells in vitro and in drug-treated mice for: HSP70, mitochondrial HSP70, HSP60, HSP56, HSP40, HSP10, and cyclophilin A. Prolonged 'rafenib/sildenafil treatment killed tumor cells and also rapidly decreased the expression of: the drug efflux pumps ABCB1 and ABCG2; and NPC1 and NTCP, receptors for Ebola/Hepatitis A and B viruses, respectively. Pre-treatment with the 'Rafenib/sildenafil combination reduced expression of the Coxsackie and Adenovirus receptor in parallel with it also reducing the ability of a serotype 5 Adenovirus or Coxsackie virus B4 to infect and to reproduce. Sorafenib/pazopanib and sildenafil was much more potent than sorafenib/pazopanib as single agents at preventing Adenovirus, Mumps, Chikungunya, Dengue, Rabies, West Nile, Yellow Fever, and Enterovirus 71 infection and reproduction. 'Rafenib drugs/pazopanib as single agents killed laboratory generated antibiotic resistant E. coli which was associated with reduced Dna K and Rec A expression. Marginally toxic doses of 'Rafenib drugs/pazopanib restored antibiotic sensitivity in pan-antibiotic resistant bacteria including multiple strains of blakpc Klebsiella pneumoniae. Thus, Dna K is an antibiotic target for sorafenib, and inhibition of GRP78/Dna K has therapeutic utility for cancer and for bacterial and viral infections.
© 2015 Wiley Periodicals, Inc.
Figures

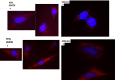
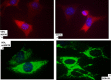
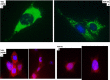
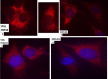







Similar articles
-
GRP78/BiP/HSPA5/Dna K is a universal therapeutic target for human disease.J Cell Physiol. 2015 Jul;230(7):1661-76. doi: 10.1002/jcp.24919. J Cell Physiol. 2015. PMID: 25546329 Free PMC article.
-
Multi-kinase inhibitors can associate with heat shock proteins through their NH2-termini by which they suppress chaperone function.Oncotarget. 2016 Mar 15;7(11):12975-96. doi: 10.18632/oncotarget.7349. Oncotarget. 2016. PMID: 26887051 Free PMC article.
-
Nexavar/Stivarga and viagra interact to kill tumor cells.J Cell Physiol. 2015 Sep;230(9):2281-98. doi: 10.1002/jcp.24961. J Cell Physiol. 2015. PMID: 25704960 Free PMC article.
-
Sorafenib.Recent Results Cancer Res. 2010;184:61-70. doi: 10.1007/978-3-642-01222-8_5. Recent Results Cancer Res. 2010. PMID: 20072831 Review.
-
Sorafenib and thyroid cancer.BioDrugs. 2013 Dec;27(6):615-28. doi: 10.1007/s40259-013-0049-y. BioDrugs. 2013. PMID: 23818056 Review.
Cited by
-
AR12 (OSU-03012) suppresses GRP78 expression and inhibits SARS-CoV-2 replication.Biochem Pharmacol. 2020 Dec;182:114227. doi: 10.1016/j.bcp.2020.114227. Epub 2020 Sep 20. Biochem Pharmacol. 2020. PMID: 32966814 Free PMC article.
-
Kinase inhibitors: look beyond the label on the bottle.Cancer Drug Resist. 2019 Dec 19;2(4):1032-1043. doi: 10.20517/cdr.2019.80. eCollection 2019. Cancer Drug Resist. 2019. PMID: 35582272 Free PMC article. Review.
-
COVID-19, Obesity, and GRP78: Unraveling the Pathological Link.J Obes Metab Syndr. 2023 Sep 30;32(3):183-196. doi: 10.7570/jomes23053. J Obes Metab Syndr. 2023. PMID: 37752707 Free PMC article. Review.
-
RNAi-based small molecule repositioning reveals clinically approved urea-based kinase inhibitors as broadly active antivirals.PLoS Pathog. 2019 Mar 18;15(3):e1007601. doi: 10.1371/journal.ppat.1007601. eCollection 2019 Mar. PLoS Pathog. 2019. PMID: 30883607 Free PMC article.
-
Synthesis and pharmacological evaluation of 1,3-diaryl substituted pyrazole based (thio)urea derivatives as potent antimicrobial agents against multi-drug resistant Staphylococcus aureus and Mycobacterium tuberculosis.RSC Med Chem. 2023 May 10;14(7):1296-1308. doi: 10.1039/d3md00079f. eCollection 2023 Jul 20. RSC Med Chem. 2023. PMID: 37484564 Free PMC article.
References
-
- Anderson K, Stott EJ, Wertz GW. 1992. Intracellular processing of the human respiratory syncytial virus fusion glycoprotein: amino acid substitutions affecting folding, transport and cleavage. J Gen Virol 73:1177–1188. - PubMed
-
- Bai LY, Weng JR, Tsai CH, Sargeant A, Lin CW, Chiu CF. 2010. OSU‐03012 sensitizes TIB‐196 myeloma cells to imatinib mesylate via AMP‐activated protein kinase and STAT3 pathways. Leuk Res 2010:816–820. - PubMed
-
- Bolt G. 2001. The measles virus (MV) glycoproteins interact with cellular chaperones in the endoplasmic reticulum and MV infection upregulates chaperone expression. Arch Virol 146:2055–2068. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

