A transcriptomic-phylogenomic analysis of the evolutionary relationships of flatworms
- PMID: 25866392
- PMCID: PMC4446793
- DOI: 10.1016/j.cub.2015.03.034
A transcriptomic-phylogenomic analysis of the evolutionary relationships of flatworms
Abstract
The interrelationships of the flatworms (phylum Platyhelminthes) are poorly resolved despite decades of morphological and molecular phylogenetic studies. The earliest-branching clades (Catenulida, Macrostomorpha, and Polycladida) share spiral cleavage and entolecithal eggs with other lophotrochozoans. Lecithoepitheliata have primitive spiral cleavage but derived ectolecithal eggs. Other orders (Rhabdocoela, Proseriata, Tricladida and relatives, and Bothrioplanida) all have derived ectolecithal eggs but have uncertain affinities to one another. The orders of parasitic Neodermata emerge from an uncertain position from within these ectolecithal classes. To tackle these problems, we have sequenced transcriptomes from 18 flatworms and 5 other metazoan groups. The addition of published data produces an alignment of >107,000 amino acids with less than 28% missing data from 27 flatworm taxa in 11 orders covering all major clades. Our phylogenetic analyses show that Platyhelminthes consist of the two clades Catenulida and Rhabditophora. Within Rhabditophora, we show the earliest-emerging branch is Macrostomorpha, not Polycladida. We show Lecithoepitheliata are not members of Neoophora but are sister group of Polycladida, implying independent origins of the ectolecithal eggs found in Lecithoepitheliata and Neoophora. We resolve Rhabdocoela as the most basally branching euneoophoran taxon. Tricladida, Bothrioplanida, and Neodermata constitute a group that appears to have lost both spiral cleavage and centrosomes. We identify Bothrioplanida as the long-sought closest free-living sister group of the parasitic Neodermata. Among parasitic orders, we show that Cestoda are closer to Trematoda than to Monogenea, rejecting the concept of the Cercomeromorpha. Our results have important implications for understanding the evolution of this major phylum.
Copyright © 2015 The Authors. Published by Elsevier Ltd.. All rights reserved.
Figures
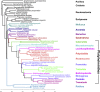
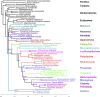


References
-
- Ehlers U. First Edition. Gustav Fischer Verlag; 1985. Das Phylogenetische System der Plathelminthes.
-
- Laumer C.E., Giribet G. Inclusive taxon sampling suggests a single, stepwise origin of ectolecithality in Platyhelminthes. Biol. J. Linn. Soc. Lond. 2014;111:570–588.
-
- Lartillot N., Lepage T., Blanquart S. PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating. Bioinformatics. 2009;25:2286–2288. - PubMed
-
- Guindon S., Dufayard J.F., Lefort V., Anisimova M., Hordijk W., Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 2010;59:307–321. - PubMed
Publication types
MeSH terms
Associated data
- SRA/SRX871300
- SRA/SRX871445
- SRA/SRX871508
- SRA/SRX871533
- SRA/SRX872314
- SRA/SRX872321
- SRA/SRX872327
- SRA/SRX872347
- SRA/SRX872356
- SRA/SRX872362
- SRA/SRX872365
- SRA/SRX872398
- SRA/SRX872402
- SRA/SRX872403
- SRA/SRX872404
- SRA/SRX872410
- SRA/SRX872414
- SRA/SRX872416
- SRA/SRX874324
- SRA/SRX875739
- SRA/SRX875742
- SRA/SRX875881
- SRA/SRX879690
- SRA/SRX883021
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

