Caspase-dependent Proteolysis of Human Ribonucleotide Reductase Small Subunits R2 and p53R2 during Apoptosis
- PMID: 25878246
- PMCID: PMC4447979
- DOI: 10.1074/jbc.M115.649640
Caspase-dependent Proteolysis of Human Ribonucleotide Reductase Small Subunits R2 and p53R2 during Apoptosis
Abstract
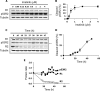
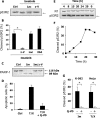
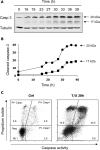
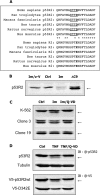
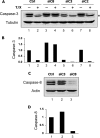
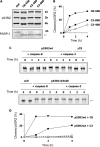
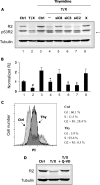
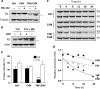
Ribonucleotide reductase (RnR) is a key enzyme synthesizing deoxyribonucleotides for DNA replication and repair. In mammals, the R1 catalytic subunit forms an active complex with either one of the two small subunits R2 and p53R2. Expression of R2 is S phase-specific and required for DNA replication. The p53R2 protein is expressed throughout the cell cycle and in quiescent cells where it provides dNTPs for mitochondrial DNA synthesis. Participation of R2 and p53R2 in DNA repair has also been suggested. In this study, we investigated the fate of the RnR subunits during apoptosis. The p53R2 protein was cleaved in a caspase-dependent manner in K-562 cells treated with inhibitors of the Bcr-Abl oncogenic kinase and in HeLa 229 cells incubated with TNF-α and cycloheximide. The cleavage site was mapped between Asp(342) and Asn(343). Caspase attack released a C-terminal p53R2 peptide of nine residues containing the conserved heptapeptide essential for R1 binding. As a consequence, the cleaved p53R2 protein was inactive. In vitro, purified caspase-3 and -8 could release the C-terminal tail of p53R2. Knocking down these caspases, but not caspase-2, -7, and -10, also inhibited p53R2 cleavage in cells committed to die via the extrinsic death receptor pathway. The R2 subunit was subjected to caspase- and proteasome-dependent proteolysis, which was prevented by siRNA targeting caspase-8. Knocking down caspase-3 was ineffective. Protein R1 was not subjected to degradation. Adding deoxyribonucleosides to restore dNTP pools transiently protected cells from apoptosis. These data identify RnR activity as a prosurvival function inactivated by proteolysis during apoptosis.
Keywords: apoptosis; caspase; nucleoside/nucleotide biosynthesis; protein degradation; ribonucleotide reductase.
© 2015 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures

Similar articles
-
Regulation of mammalian ribonucleotide reduction and dNTP pools after DNA damage and in resting cells.J Biol Chem. 2006 Mar 24;281(12):7834-41. doi: 10.1074/jbc.M512894200. Epub 2006 Jan 24. J Biol Chem. 2006. PMID: 16436374
-
Deoxyribonucleotide metabolism in cycling and resting human fibroblasts with a missense mutation in p53R2, a subunit of ribonucleotide reductase.J Biol Chem. 2011 Apr 1;286(13):11132-40. doi: 10.1074/jbc.M110.202283. Epub 2011 Feb 5. J Biol Chem. 2011. PMID: 21297166 Free PMC article.
-
Stable suppression of the R2 subunit of ribonucleotide reductase by R2-targeted short interference RNA sensitizes p53(-/-) HCT-116 colon cancer cells to DNA-damaging agents and ribonucleotide reductase inhibitors.J Biol Chem. 2004 Jun 25;279(26):27030-8. doi: 10.1074/jbc.M402056200. Epub 2004 Apr 19. J Biol Chem. 2004. PMID: 15096505
-
Cellular regulation of ribonucleotide reductase in eukaryotes.Semin Cell Dev Biol. 2014 Jun;30:97-103. doi: 10.1016/j.semcdb.2014.03.030. Epub 2014 Apr 2. Semin Cell Dev Biol. 2014. PMID: 24704278 Review.
-
Akt and p53R2, partners that dictate the progression and invasiveness of cancer.DNA Repair (Amst). 2014 Oct;22:24-9. doi: 10.1016/j.dnarep.2014.07.001. Epub 2014 Aug 1. DNA Repair (Amst). 2014. PMID: 25086499 Review.
Cited by
-
Still no Rest for the Reductases: Ribonucleotide Reductase (RNR) Structure and Function: An Update.Subcell Biochem. 2022;99:155-197. doi: 10.1007/978-3-031-00793-4_5. Subcell Biochem. 2022. PMID: 36151376 Review.
-
Folic acid tagged nanoceria as a novel therapeutic agent in ovarian cancer.BMC Cancer. 2016 Mar 15;16:220. doi: 10.1186/s12885-016-2206-4. BMC Cancer. 2016. PMID: 26979107 Free PMC article.
-
Proteome profiling of different rat brain regions reveals the modulatory effect of prolonged maternal separation on proteins involved in cell death-related processes.Biol Res. 2021 Feb 8;54(1):4. doi: 10.1186/s40659-021-00327-5. Biol Res. 2021. PMID: 33557947 Free PMC article.
-
RRM2B Is Frequently Amplified Across Multiple Tumor Types: Implications for DNA Repair, Cellular Survival, and Cancer Therapy.Front Genet. 2021 Mar 12;12:628758. doi: 10.3389/fgene.2021.628758. eCollection 2021. Front Genet. 2021. PMID: 33868369 Free PMC article.
References
-
- Nordlund P., Reichard P. (2006) Ribonucleotide reductases. Annu. Rev. Biochem. 75, 681–706 - PubMed
-
- Pontarin G., Ferraro P., Håkansson P., Thelander L., Reichard P., Bianchi V. (2007) p53R2-dependent ribonucleotide reduction provides deoxyribonucleotides in quiescent human fibroblasts in the absence of induced DNA damage. J. Biol. Chem. 282, 16820–16828 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

