Comparative transcriptomic analysis reveals similarities and dissimilarities in Saccharomyces cerevisiae wine strains response to nitrogen availability
- PMID: 25884705
- PMCID: PMC4401569
- DOI: 10.1371/journal.pone.0122709
Comparative transcriptomic analysis reveals similarities and dissimilarities in Saccharomyces cerevisiae wine strains response to nitrogen availability
Abstract
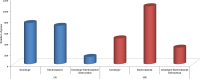
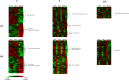
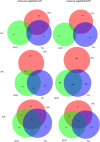
Nitrogen levels in grape-juices are of major importance in winemaking ensuring adequate yeast growth and fermentation performance. Here we used a comparative transcriptome analysis to uncover wine yeasts responses to nitrogen availability during fermentation. Gene expression was assessed in three genetically and phenotypically divergent commercial wine strains (CEG, VL1 and QA23), under low (67 mg/L) and high nitrogen (670 mg/L) regimes, at three time points during fermentation (12 h, 24 h and 96 h). Two-way ANOVA analysis of each fermentation condition led to the identification of genes whose expression was dependent on strain, fermentation stage and on the interaction of both factors. The high fermenter yeast strain QA23 was more clearly distinct from the other two strains, by differential expression of genes involved in flocculation, mitochondrial functions, energy generation and protein folding and stabilization. For all strains, higher transcriptional variability due to fermentation stage was seen in the high nitrogen fermentations. A positive correlation between maximum fermentation rate and the expression of genes involved in stress response was observed. The finding of common genes correlated with both fermentation activity and nitrogen up-take underlies the role of nitrogen on yeast fermentative fitness. The comparative analysis of genes differentially expressed between both fermentation conditions at 12 h, where the main difference was the level of nitrogen available, showed the highest variability amongst strains revealing strain-specific responses. Nevertheless, we were able to identify a small set of genes whose expression profiles can quantitatively assess the common response of the yeast strains to varying nitrogen conditions. The use of three contrasting yeast strains in gene expression analysis prompts the identification of more reliable, accurate and reproducible biomarkers that will facilitate the diagnosis of deficiency of this nutrient in the grape-musts and the development of strategies to optimize yeast performance in industrial fermentations.
Conflict of interest statement
Figures

Similar articles
-
The role of nitrogen uptake on the competition ability of three vineyard Saccharomyces cerevisiae strains.Int J Food Microbiol. 2017 Oct 3;258:1-11. doi: 10.1016/j.ijfoodmicro.2017.07.006. Epub 2017 Jul 13. Int J Food Microbiol. 2017. PMID: 28735228
-
Use of commercial or indigenous yeast impacts the S. cerevisiae transcriptome during wine fermentation.Microbiol Spectr. 2024 Nov 5;12(11):e0119424. doi: 10.1128/spectrum.01194-24. Epub 2024 Sep 17. Microbiol Spectr. 2024. PMID: 39287451 Free PMC article.
-
Optogenetic control of horizontally acquired genes prevent stuck fermentations in yeast.Microbiol Spectr. 2025 Feb 4;13(2):e0179424. doi: 10.1128/spectrum.01794-24. Epub 2025 Jan 8. Microbiol Spectr. 2025. PMID: 39772912 Free PMC article.
-
Disentangling the genetic bases of Saccharomyces cerevisiae nitrogen consumption and adaptation to low nitrogen environments in wine fermentation.Biol Res. 2020 Jan 9;53(1):2. doi: 10.1186/s40659-019-0270-3. Biol Res. 2020. PMID: 31918759 Free PMC article. Review.
-
Responses of Saccharomyces cerevisiae to nitrogen starvation in wine alcoholic fermentation.Appl Microbiol Biotechnol. 2015 Sep;99(17):7025-34. doi: 10.1007/s00253-015-6810-z. Epub 2015 Jul 23. Appl Microbiol Biotechnol. 2015. PMID: 26201494 Review.
Cited by
-
Differential Gene Expression and Allele Frequency Changes Favour Adaptation of a Heterogeneous Yeast Population to Nitrogen-Limited Fermentations.Front Microbiol. 2020 Jun 15;11:1204. doi: 10.3389/fmicb.2020.01204. eCollection 2020. Front Microbiol. 2020. PMID: 32612585 Free PMC article.
-
Secondary Aroma: Influence of Wine Microorganisms in Their Aroma Profile.Foods. 2020 Dec 27;10(1):51. doi: 10.3390/foods10010051. Foods. 2020. PMID: 33375439 Free PMC article. Review.
-
In Vivo Analysis of NH4+ Transport and Central Nitrogen Metabolism in Saccharomyces cerevisiae during Aerobic Nitrogen-Limited Growth.Appl Environ Microbiol. 2016 Dec 1;82(23):6831-6845. doi: 10.1128/AEM.01547-16. Epub 2016 Sep 16. Appl Environ Microbiol. 2016. PMID: 27637876 Free PMC article.
-
The Timing of Nitrogen Addition Impacts Yeast Genes Expression and the Production of Aroma Compounds During Wine Fermentation.Front Microbiol. 2022 Feb 22;13:829786. doi: 10.3389/fmicb.2022.829786. eCollection 2022. Front Microbiol. 2022. PMID: 35273585 Free PMC article.
-
Pesticides, cosmetics, drugs: identical and opposite influences of various molecular features as measures of endpoints similarity and dissimilarity.Mol Divers. 2021 May;25(2):1137-1144. doi: 10.1007/s11030-020-10085-3. Epub 2020 Apr 23. Mol Divers. 2021. PMID: 32323128
References
-
- Pretorius IS. Tailoring wine yeast for the new millennium: novel approaches to the ancient art of winemaking. Yeast. 2000; 16:675–729. - PubMed
-
- Henschke PA, Jiranek V. Yeasts-Metabolism of nitrogen compounds In: Fleet GH, editor. Wine Microbiology and Biotechnology. Harwood Academic, Switzerland; 1993. pp 77–164.
-
- Bisson LF, Butzke CE. Diagnosis and rectification of stuck and sluggish fermentations. Am J Enol Vitic. 2000; 51: 168–177.
-
- Bezenger MC, Navarro JM. Alcoholic fermentation: Model accounting for initial nitrogen influence. Biotechnol and Bioeng. 1988; 31: 747–9. - PubMed
-
- Bisson LF. Stuck and sluggish fermentations. Am J Enol Vitic. 1999; 50:107–119.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

