Comparison of the three-dimensional organization of sperm and fibroblast genomes using the Hi-C approach
- PMID: 25886366
- PMCID: PMC4434584
- DOI: 10.1186/s13059-015-0642-0
Comparison of the three-dimensional organization of sperm and fibroblast genomes using the Hi-C approach
Erratum in
-
Erratum to: comparison of the three-dimensional organization of sperm and fibroblast genomes using the Hi-C approach.Genome Biol. 2016 Jan 14;17:6. doi: 10.1186/s13059-016-0868-5. Genome Biol. 2016. PMID: 26769583 Free PMC article. No abstract available.
Abstract
Background: The three-dimensional organization of the genome is tightly connected to its biological function. The Hi-C approach was recently introduced as a method that can be used to identify higher-order chromatin interactions genome-wide. The aim of this study was to determine genome-wide chromatin interaction frequencies using the Hi-C approach in mouse sperm cells and embryonic fibroblasts.
Results: The obtained data demonstrate that the three-dimensional genome organizations of sperm and fibroblast cells show a high degree of similarity both with each other and with the previously described mouse embryonic stem cells. Both A- and B-compartments and topologically associated domains are present in spermatozoa and fibroblasts. Nevertheless, sperm cells and fibroblasts exhibit statistically significant differences between each other in the contact probabilities of defined loci. Tight packaging of the sperm genome results in an enrichment of long-range contacts compared with the fibroblasts. However, only 30% of the differences in the number of contacts are based on differences in the densities of their genome packages; the main source of the differences is the gain or loss of contacts that are specific for defined genome regions. We find that the dependence of the contact probability on genomic distance for sperm is close to the dependence predicted for the fractal globular folding of chromatin.
Conclusions: Overall, we can conclude that the three-dimensional structure of the genome is passed through generations without being dramatically changed in sperm cells.
Figures
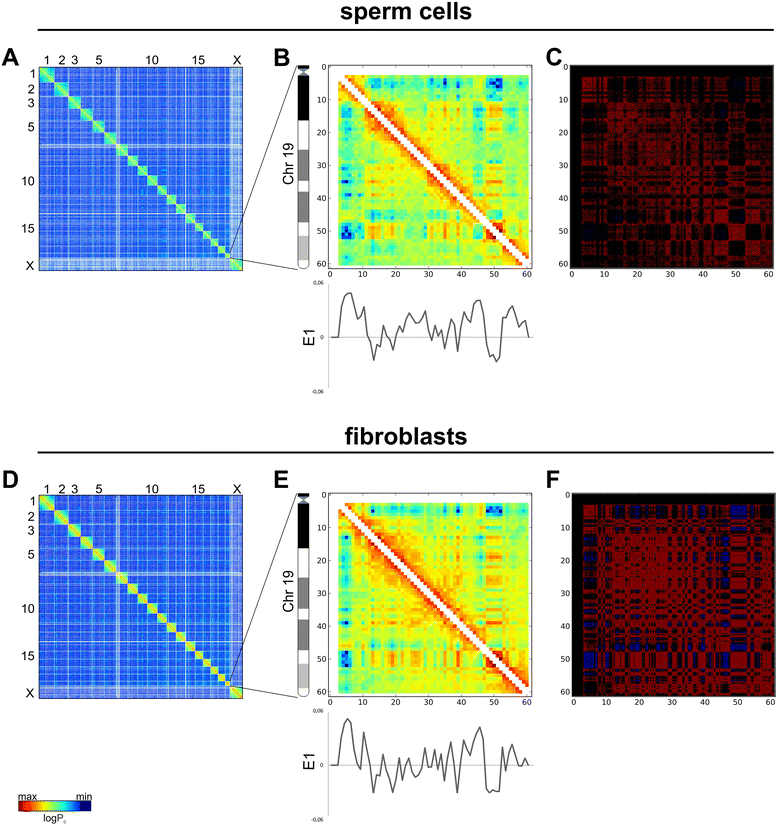





References
-
- Markaki Y, Gunkel M, Schermelleh L, Beichmanis S, Neumann J, Heidemann M, et al. Functional nuclear organization of transcription and DNA replication: a topographical marriage between chromatin domains and the interchromatin compartment. Cold Spring Harb Symp Quant Biol. 2010;75:475–92. doi: 10.1101/sqb.2010.75.042. - DOI - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources

