Molecular epidemiological study of feline coronavirus strains in Japan using RT-PCR targeting nsp14 gene
- PMID: 25889235
- PMCID: PMC4359392
- DOI: 10.1186/s12917-015-0372-2
Molecular epidemiological study of feline coronavirus strains in Japan using RT-PCR targeting nsp14 gene
Abstract
Background: Feline infectious peritonitis is a fatal disease of cats caused by infection with feline coronavirus (FCoV). For detecting or genotyping of FCoV, some RT-PCR plus nested PCR techniques have been reported previously. However, referring to the whole genome sequences (WGSs) registered at NCBI, there are no detection methods that can tolerate the genetic diversity among FCoV population. In addition, the quasispecies nature of FCoV, which consists of heterogeneous variants, has been also demonstrated; thus, a universal method for heteropopulations of FCoV variants in clinical specimens is desirable.
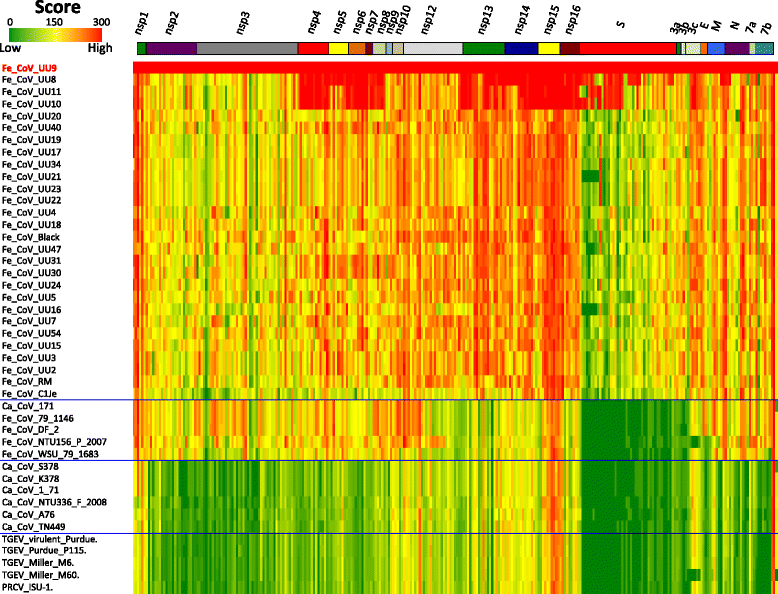
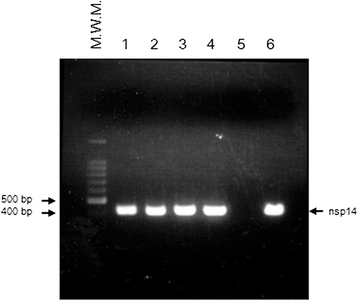
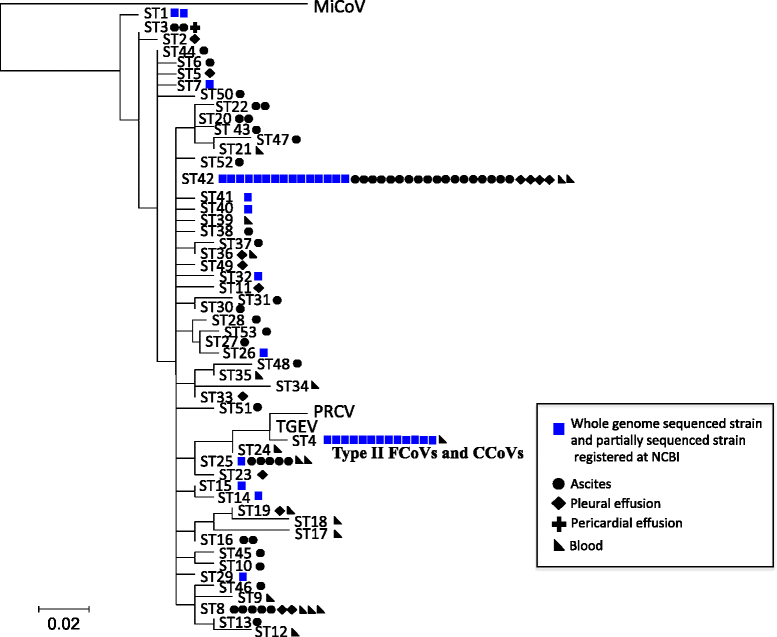
Results: To develop an RT-PCR method for detection and genotyping of FCoV, we performed comparative genome analysis using WGSs of 32 FCoV, 7 CCoV and 5 TGEV strains obtained from NCBI. As the PCR target, we focused on the nsp14 coding region, which is highly conserved and phylogenetically informative, and developed a PCR method targeting nsp14 partial sequences. Among 103 ascites, 45 pleural effusion and 214 blood specimens from clinically ill cats, we could detect FCoV in 55 (53.4%), 14 (31.1%) and 19 (8.9%) specimens using the present method. Direct sequencing of PCR products and phylogenetic analysis allowed discrimination between type I- and II-FCoV serotypes. Our nsp14 amino acid sequence typing (nsp14 aa ST) showed that the FCoV clone with sequence type (ST) 42, which was the most predominant genotype of WGS strains, was prevalent in domestic cats in Japan.
Conclusions: Our nsp14 PCR scheme will contribute to virus detection, epidemiology and ecology of FCoV strains.
Figures

Similar articles
-
Circulation and genetic diversity of Feline coronavirus type I and II from clinically healthy and FIP-suspected cats in China.Transbound Emerg Dis. 2019 Mar;66(2):763-775. doi: 10.1111/tbed.13081. Epub 2018 Dec 5. Transbound Emerg Dis. 2019. PMID: 30468573 Free PMC article.
-
Sensitivity and specificity of a real-time reverse transcriptase polymerase chain reaction detecting feline coronavirus mutations in effusion and serum/plasma of cats to diagnose feline infectious peritonitis.BMC Vet Res. 2017 Aug 2;13(1):228. doi: 10.1186/s12917-017-1147-8. BMC Vet Res. 2017. PMID: 28768514 Free PMC article.
-
Comparative sequence analysis of the accessory and nucleocapsid genes of feline coronavirus strains isolated from cats diagnosed with effusive feline infectious peritonitis.Arch Virol. 2021 Oct;166(10):2779-2787. doi: 10.1007/s00705-021-05188-7. Epub 2021 Aug 7. Arch Virol. 2021. PMID: 34363535 Free PMC article.
-
A Tale of Two Viruses: The Distinct Spike Glycoproteins of Feline Coronaviruses.Viruses. 2020 Jan 10;12(1):83. doi: 10.3390/v12010083. Viruses. 2020. PMID: 31936749 Free PMC article. Review.
-
Feline infectious peritonitis: still an enigma?Vet Pathol. 2014 Mar;51(2):505-26. doi: 10.1177/0300985814522077. Vet Pathol. 2014. PMID: 24569616 Review.
Cited by
-
Near-complete genome sequencing of the feline coronavirus serotype I strain FIPV-Aqua from a cat with feline infectious peritonitis in Japan.Microbiol Resour Announc. 2025 Jun 12;14(6):e0035725. doi: 10.1128/mra.00357-25. Epub 2025 May 20. Microbiol Resour Announc. 2025. PMID: 40391907 Free PMC article.
-
Whole genome characterization of feline coronaviruses in Thailand: evidence of genetic recombination and mutation M1058L in pathotype switch.Front Vet Sci. 2025 Feb 14;12:1451967. doi: 10.3389/fvets.2025.1451967. eCollection 2025. Front Vet Sci. 2025. PMID: 40027354 Free PMC article.
-
Genetic and Phylogenetic Analysis of Feline Coronavirus in Guangxi Province of China from 2021 to 2024.Vet Sci. 2024 Sep 25;11(10):455. doi: 10.3390/vetsci11100455. Vet Sci. 2024. PMID: 39453047 Free PMC article.
-
Epidemiology and Comparative Analyses of the S Gene on Feline Coronavirus in Central China.Pathogens. 2022 Apr 12;11(4):460. doi: 10.3390/pathogens11040460. Pathogens. 2022. PMID: 35456135 Free PMC article.
-
Characterization of antiviral T cell responses during primary and secondary challenge of laboratory cats with feline infectious peritonitis virus (FIPV).BMC Vet Res. 2019 May 22;15(1):165. doi: 10.1186/s12917-019-1909-6. BMC Vet Res. 2019. PMID: 31118053 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

