Unveiling the roles of HBV polymerase for new antiviral strategies
- PMID: 25893003
- PMCID: PMC4399241
- DOI: 10.2217/fvl.14.113
Unveiling the roles of HBV polymerase for new antiviral strategies
Abstract
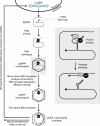
Infection with HBV is common worldwide, with over 350 million chronic carriers. Chronic HBV infection is associated with cirrhosis and hepatocellular carcinoma. All currently available oral antivirals are directed against the HBV polymerase enzyme, a reverse transcriptase. HBV polymerase contains several important domains and motifs which define its functions and reveal ways to further target it. This enzyme executes many functions required for the HBV replication cycle, including viral RNA binding, RNA packaging, protein priming, template switching, DNA synthesis and RNA degradation. In addition, HBV polymerase must interact with host proteins for its functions. Future therapeutics may inhibit not only the DNA synthesis steps which are carried out by the reverse transcriptase domain (as all current antivirals do) but other domains, functions and interactions which are essential to the HBV replication cycle.
Keywords: HBV; antiviral targets; drug resistance; replication cycle; reverse transcriptase.
Figures


Similar articles
-
Hepatitis B virus reverse transcriptase - Target of current antiviral therapy and future drug development.Antiviral Res. 2015 Nov;123:132-7. doi: 10.1016/j.antiviral.2015.09.011. Epub 2015 Sep 25. Antiviral Res. 2015. PMID: 26408354 Free PMC article. Review.
-
RNA-Binding Motif Protein 24 (RBM24) Is Involved in Pregenomic RNA Packaging by Mediating Interaction between Hepatitis B Virus Polymerase and the Epsilon Element.J Virol. 2019 Mar 5;93(6):e02161-18. doi: 10.1128/JVI.02161-18. Print 2019 Mar 15. J Virol. 2019. PMID: 30626666 Free PMC article.
-
Hepatitis B virus reverse transcriptase: diverse functions as classical and emerging targets for antiviral intervention.Emerg Microbes Infect. 2013 Sep;2(9):e56. doi: 10.1038/emi.2013.56. Epub 2013 Sep 4. Emerg Microbes Infect. 2013. PMID: 26038488 Free PMC article. Review.
-
Mapping of Functional Subdomains in the Terminal Protein Domain of Hepatitis B Virus Polymerase.J Virol. 2017 Jan 18;91(3):e01785-16. doi: 10.1128/JVI.01785-16. Print 2017 Feb 1. J Virol. 2017. PMID: 27852858 Free PMC article.
-
Treatment for hepatitis B in patients with drug resistance.Ann Transl Med. 2016 Sep;4(18):334. doi: 10.21037/atm.2016.09.19. Ann Transl Med. 2016. PMID: 27761438 Free PMC article. Review.
Cited by
-
Discovery and Selection of Hepatitis B Virus-Derived T Cell Epitopes for Global Immunotherapy Based on Viral Indispensability, Conservation, and HLA-Binding Strength.J Virol. 2020 Mar 17;94(7):e01663-19. doi: 10.1128/JVI.01663-19. Print 2020 Mar 17. J Virol. 2020. PMID: 31852786 Free PMC article.
-
3,7-Dihydroxytropolones Inhibit Initiation of Hepatitis B Virus Minus-Strand DNA Synthesis.Molecules. 2020 Sep 27;25(19):4434. doi: 10.3390/molecules25194434. Molecules. 2020. PMID: 32992516 Free PMC article.
-
Identification of novel hepatitis B virus therapeutic vaccine candidates derived from polymerase protein.Aging (Albany NY). 2021 May 20;13(10):14372-14384. doi: 10.18632/aging.203053. Epub 2021 May 20. Aging (Albany NY). 2021. PMID: 34016795 Free PMC article.
-
Viral and cellular oncogenes promote immune evasion.Oncogene. 2022 Feb;41(7):921-929. doi: 10.1038/s41388-021-02145-1. Epub 2022 Jan 13. Oncogene. 2022. PMID: 35022539 Free PMC article. Review.
-
Viral hepatitis: Milestones, unresolved issues, and future goals.World J Gastroenterol. 2021 Jul 28;27(28):4603-4638. doi: 10.3748/wjg.v27.i28.4603. World J Gastroenterol. 2021. PMID: 34366625 Free PMC article. Review.
References
-
- Summers J, Mason WS. Replication of the genome of a hepatitis B – like virus by reverse transcription of an RNA intermediate. Cell. 1982;29(2):403–415. - PubMed
-
- Bond W, Favero M, Petersen N, Gravelle C, Ebert J, Maynard J. Survival of hepatitis B virus after drying and storage for one week. Lancet. 1981;317(8219):550–551. - PubMed
-
- McMahon BJ, Alward WLM, Hall DB, et al. Acute hepatitis B virus infection: relation of age to the clinical expression of disease and subsequent development of the carrier state. J. Infect. Dis. 1985;151(4):599–603. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
