Serratia marcescens Cyclic AMP Receptor Protein Controls Transcription of EepR, a Novel Regulator of Antimicrobial Secondary Metabolites
- PMID: 25897029
- PMCID: PMC4518835
- DOI: 10.1128/JB.00136-15
Serratia marcescens Cyclic AMP Receptor Protein Controls Transcription of EepR, a Novel Regulator of Antimicrobial Secondary Metabolites
Abstract
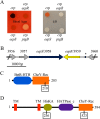
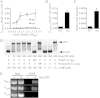
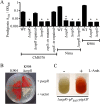
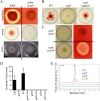
Serratia marcescens generates secondary metabolites and secreted enzymes, and it causes hospital infections and community-acquired ocular infections. Previous studies identified cyclic AMP (cAMP) receptor protein (CRP) as an indirect inhibitor of antimicrobial secondary metabolites. Here, we identified a putative two-component regulator that suppressed crp mutant phenotypes. Evidence supports that the putative response regulator eepR was directly transcriptionally inhibited by cAMP-CRP. EepR and the putative sensor kinase EepS were necessary for the biosynthesis of secondary metabolites, including prodigiosin- and serratamolide-dependent phenotypes, swarming motility, and hemolysis. Recombinant EepR bound to the prodigiosin and serratamolide promoters in vitro. Together, these data introduce a novel regulator of secondary metabolites that directly connects the broadly conserved metabolism regulator CRP with biosynthetic genes that may contribute to competition with other microbes.
Importance: This study identifies a new transcription factor that is directly controlled by a broadly conserved transcription factor, CRP. CRP is well studied in its role to help bacteria respond to the amount of nutrients in their environment. The new transcription factor EepR is essential for the bacterium Serratia marcescens to produce two biologically active compounds, prodigiosin and serratamolide. These two compounds are antimicrobial and may allow S. marcescens to compete for limited nutrients with other microorganisms. Results from this study tie together the CRP environmental nutrient sensor with a new regulator of antimicrobial compounds. Beyond microbial ecology, prodigiosin and serratamolide have therapeutic potential; therefore, understanding their regulation is important for both applied and basic science.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures

Similar articles
-
A Serratia marcescens PigP homolog controls prodigiosin biosynthesis, swarming motility and hemolysis and is regulated by cAMP-CRP and HexS.PLoS One. 2013;8(3):e57634. doi: 10.1371/journal.pone.0057634. Epub 2013 Mar 1. PLoS One. 2013. PMID: 23469212 Free PMC article.
-
Suppressor analysis of eepR mutant defects reveals coordinate regulation of secondary metabolites and serralysin biosynthesis by EepR and HexS.Microbiology (Reading). 2017 Feb;163(2):280-288. doi: 10.1099/mic.0.000422. Microbiology (Reading). 2017. PMID: 28270264 Free PMC article.
-
An IgaA/UmoB Family Protein from Serratia marcescens Regulates Motility, Capsular Polysaccharide Biosynthesis, and Secondary Metabolite Production.Appl Environ Microbiol. 2018 Mar 1;84(6):e02575-17. doi: 10.1128/AEM.02575-17. Print 2018 Mar 15. Appl Environ Microbiol. 2018. PMID: 29305504 Free PMC article.
-
Syn, anti, and finally both conformations of cyclic AMP are involved in the CRP-dependent transcription initiation mechanism in E. coli lac operon.Cell Biochem Funct. 2008 Jun;26(4):399-405. doi: 10.1002/cbf.1462. Cell Biochem Funct. 2008. PMID: 18338329 Review.
-
Recent advancements in high-level synthesis of the promising clinical drug, prodigiosin.Appl Microbiol Biotechnol. 2019 Feb;103(4):1667-1680. doi: 10.1007/s00253-018-09611-z. Epub 2019 Jan 12. Appl Microbiol Biotechnol. 2019. PMID: 30637495 Review.
Cited by
-
The Rcs Stress Response System Regulator GumB Modulates Serratia marcescens-Induced Inflammation and Bacterial Proliferation in a Rabbit Keratitis Model and Cytotoxicity In Vitro.Infect Immun. 2021 Jul 15;89(8):e0011121. doi: 10.1128/IAI.00111-21. Epub 2021 Jul 15. Infect Immun. 2021. PMID: 33820815 Free PMC article.
-
Exploitation of a "hockey-puck" phenotype to identify pilus and biofilm regulators in Serratia marcescens through genetic analysis.Can J Microbiol. 2016 Jan;62(1):83-93. doi: 10.1139/cjm-2015-0566. Epub 2015 Nov 3. Can J Microbiol. 2016. PMID: 26640000 Free PMC article.
-
Transcriptional factor OmpR positively regulates prodigiosin biosynthesis in Serratia marcescens FZSF02 by binding with the promoter of the prodigiosin cluster.Front Microbiol. 2022 Nov 17;13:1041146. doi: 10.3389/fmicb.2022.1041146. eCollection 2022. Front Microbiol. 2022. PMID: 36466667 Free PMC article.
-
Thermoregulation of Prodigiosin Biosynthesis by Serratia marcescens is Controlled at the Transcriptional Level and Requires HexS.Pol J Microbiol. 2019;68(1):43-50. doi: 10.21307/pjm-2019-005. Pol J Microbiol. 2019. PMID: 31050252 Free PMC article.
-
Multicopy expression of sigma factor RpoH reduces prodigiosin biosynthesis in Serratia marcescens FS14.Antonie Van Leeuwenhoek. 2023 Nov;116(11):1197-1208. doi: 10.1007/s10482-023-01875-4. Epub 2023 Sep 20. Antonie Van Leeuwenhoek. 2023. PMID: 37728826
References
-
- Komeda Y, Suzuki H, Ishidsu I, Iino T. 1976. The role of cAMP in flagellation of Salmonella typhimurium. Mol Gen Genet 142:289–298. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

