The BioMart community portal: an innovative alternative to large, centralized data repositories
- PMID: 25897122
- PMCID: PMC4489294
- DOI: 10.1093/nar/gkv350
The BioMart community portal: an innovative alternative to large, centralized data repositories
Abstract
The BioMart Community Portal (www.biomart.org) is a community-driven effort to provide a unified interface to biomedical databases that are distributed worldwide. The portal provides access to numerous database projects supported by 30 scientific organizations. It includes over 800 different biological datasets spanning genomics, proteomics, model organisms, cancer data, ontology information and more. All resources available through the portal are independently administered and funded by their host organizations. The BioMart data federation technology provides a unified interface to all the available data. The latest version of the portal comes with many new databases that have been created by our ever-growing community. It also comes with better support and extensibility for data analysis and visualization tools. A new addition to our toolbox, the enrichment analysis tool is now accessible through graphical and web service interface. The BioMart community portal averages over one million requests per day. Building on this level of service and the wealth of information that has become available, the BioMart Community Portal has introduced a new, more scalable and cheaper alternative to the large data stores maintained by specialized organizations.
© The Author(s) 2015. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
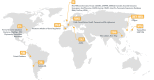


References
-
- Durinck S., Moreau Y., Kasprzyk A., Davis S., De Moor B., Brazma A., Huber W. BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics. 2005;21:3439–3440. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

