Noncognate DNA damage prevents the formation of the active conformation of the Y-family DNA polymerases DinB and DNA polymerase κ
- PMID: 25899385
- PMCID: PMC4504746
- DOI: 10.1111/febs.13304
Noncognate DNA damage prevents the formation of the active conformation of the Y-family DNA polymerases DinB and DNA polymerase κ
Abstract
Y-family DNA polymerases are specialized to copy damaged DNA, and are associated with increased mutagenesis, owing to their low fidelity. It is believed that the mechanism of nucleotide selection by Y-family DNA polymerases involves conformational changes preceding nucleotidyl transfer, but there is limited experimental evidence for such structural changes. In particular, nucleotide-induced conformational changes in bacterial or eukaryotic Y-family DNA polymerases have, to date, not been extensively characterized. Using hydrogen-deuterium exchange mass spectrometry, we demonstrate here that the Escherichia coli Y-family DNA polymerase DinB and its human ortholog DNA polymerase κ undergo a conserved nucleotide-induced conformational change in the presence of undamaged DNA and the correct incoming nucleotide. Notably, this holds true for damaged DNA containing N(2) -furfuryl-deoxyguanosine, which is efficiently copied by these two polymerases, but not for damaged DNA containing the major groove modification O(6) -methyl-deoxyguanosine, which is a poor substrate. Our observations suggest that DinB and DNA polymerase κ utilize a common mechanism for nucleotide selection involving a conserved prechemical conformational transition promoted by the correct nucleotide and only preferred DNA substrates.
Keywords: DNA replication; conformational change; hydrogen exchange mass spectrometry; nucleotide selection; substrate specificity.
© 2015 FEBS.
Conflict of interest statement
J.R.E. is a paid consultant of the Waters Corporation.
Figures



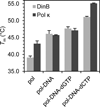


References
-
- Echols H, Goodman MF. Fidelity mechanisms in DNA replication. Annu Rev Biochem. 1991;60:477–511. - PubMed
-
- Joyce CM, Benkovic SJ. DNA polymerase fidelity: kinetics, structure, and checkpoints. Biochemistry. 2004;43:14317–14324. - PubMed
-
- Jarosz DF, Godoy VG, DeLaney JC, Essigmann JM, Walker GC. A single amino acid governs enhanced activity of DinB DNA polymerases on damaged templates. Nature. 2006;439:225–228. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

