CRISPR immunity drives rapid phage genome evolution in Streptococcus thermophilus
- PMID: 25900652
- PMCID: PMC4453577
- DOI: 10.1128/mBio.00262-15
CRISPR immunity drives rapid phage genome evolution in Streptococcus thermophilus
Abstract
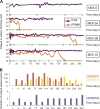
Many bacteria rely on CRISPR-Cas systems to provide adaptive immunity against phages, predation by which can shape the ecology and functioning of microbial communities. To characterize the impact of CRISPR immunization on phage genome evolution, we performed long-term bacterium-phage (Streptococcus thermophilus-phage 2972) coevolution experiments. We found that in this species, CRISPR immunity drives fixation of single nucleotide polymorphisms that accumulate exclusively in phage genome regions targeted by CRISPR. Mutation rates in phage genomes highly exceed those of the host. The presence of multiple phages increased phage persistence by enabling recombination-based formation of chimeric phage genomes in which sequences heavily targeted by CRISPR were replaced. Collectively, our results establish CRISPR-Cas adaptive immunity as a key driver of phage genome evolution under the conditions studied and highlight the importance of multiple coexisting phages for persistence in natural systems.
Importance: Phages remain an enigmatic part of the biosphere. As predators, they challenge the survival of host bacteria and archaea and set off an "arms race" involving host immunization countered by phage mutation. The CRISPR-Cas system is adaptive: by capturing fragments of a phage genome upon exposure, the host is positioned to counteract future infections. To investigate this process, we initiated massive deep-sequencing experiments with a host and infective phage and tracked the coevolution of both populations over hundreds of days. In the present study, we found that CRISPR immunity drives the accumulation of phage genome rearrangements (which enable longer phage survival) and escape mutations, establishing CRISPR as one of the fundamental drivers of phage evolution.
Copyright © 2015 Paez-Espino et al.
Figures




References
Publication types
MeSH terms
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

