Loss of the N-terminal methyltransferase NRMT1 increases sensitivity to DNA damage and promotes mammary oncogenesis
- PMID: 25909287
- PMCID: PMC4494936
- DOI: 10.18632/oncotarget.3653
Loss of the N-terminal methyltransferase NRMT1 increases sensitivity to DNA damage and promotes mammary oncogenesis
Abstract
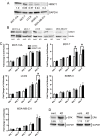
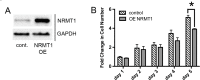
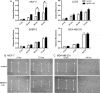
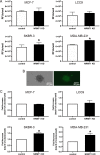
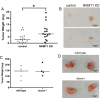
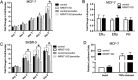
Though discovered over four decades ago, the function of N-terminal methylation has mostly remained a mystery. Our discovery of the first mammalian N-terminal methyltransferase, NRMT1, has led to the discovery of many new functions for N-terminal methylation, including regulation of DNA/protein interactions, accurate mitotic division, and nucleotide excision repair (NER). Here we test whether NRMT1 is also important for DNA double-strand break (DSB) repair, and given its previously known roles in cell cycle regulation and the DNA damage response, assay if NRMT1 is acting as a tumor suppressor. We find that NRMT1 knockdown significantly enhances the sensitivity of breast cancer cell lines to both etoposide treatment and γ-irradiation, as well as, increases proliferation rate, invasive potential, anchorage-independent growth, xenograft tumor size, and tamoxifen sensitivity. Interestingly, this positions NRMT1 as a tumor suppressor protein involved in multiple DNA repair pathways, and indicates, similar to BRCA1 and BRCA2, its loss may result in tumors with enhanced sensitivity to diverse DNA damaging chemotherapeutics.
Keywords: DNA damage; DNA repair; N-terminal methylation; NRMT1; breast cancer.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Select human cancer mutants of NRMT1 alter its catalytic activity and decrease N-terminal trimethylation.Protein Sci. 2017 Aug;26(8):1639-1652. doi: 10.1002/pro.3202. Epub 2017 Jun 11. Protein Sci. 2017. PMID: 28556566 Free PMC article.
-
NRMT2 is an N-terminal monomethylase that primes for its homologue NRMT1.Biochem J. 2013 Dec 15;456(3):453-62. doi: 10.1042/BJ20131163. Biochem J. 2013. PMID: 24090352 Free PMC article.
-
NRMT1 knockout mice exhibit phenotypes associated with impaired DNA repair and premature aging.Mech Ageing Dev. 2015 Mar;146-148:42-52. doi: 10.1016/j.mad.2015.03.012. Epub 2015 Apr 2. Mech Ageing Dev. 2015. PMID: 25843235 Free PMC article.
-
The role of the BRCA1 tumor suppressor in DNA double-strand break repair.Mol Cancer Res. 2005 Oct;3(10):531-9. doi: 10.1158/1541-7786.MCR-05-0192. Mol Cancer Res. 2005. PMID: 16254187 Review.
-
Therapeutic exploitation of tumor cell defects in homologous recombination.Anticancer Agents Med Chem. 2008 May;8(4):448-60. doi: 10.2174/187152008784220267. Anticancer Agents Med Chem. 2008. PMID: 18473729 Review.
Cited by
-
The comprehensive analysis of the prognostic and functional role of N-terminal methyltransferases 1 in pan-cancer.PeerJ. 2023 Oct 24;11:e16263. doi: 10.7717/peerj.16263. eCollection 2023. PeerJ. 2023. PMID: 37901469 Free PMC article.
-
Select human cancer mutants of NRMT1 alter its catalytic activity and decrease N-terminal trimethylation.Protein Sci. 2017 Aug;26(8):1639-1652. doi: 10.1002/pro.3202. Epub 2017 Jun 11. Protein Sci. 2017. PMID: 28556566 Free PMC article.
-
Chemical Biology of Protein N-Terminal Methyltransferases.Chembiochem. 2019 Apr 15;20(8):976-984. doi: 10.1002/cbic.201800615. Epub 2019 Feb 13. Chembiochem. 2019. PMID: 30479015 Free PMC article. Review.
-
Methyltransferases: Functions and Applications.Chembiochem. 2022 Sep 16;23(18):e202200212. doi: 10.1002/cbic.202200212. Epub 2022 Jul 5. Chembiochem. 2022. PMID: 35691829 Free PMC article. Review.
-
Chemoproteomic Study Uncovers HemK2/KMT9 As a New Target for NTMT1 Bisubstrate Inhibitors.ACS Chem Biol. 2021 Jul 16;16(7):1234-1242. doi: 10.1021/acschembio.1c00279. Epub 2021 Jun 30. ACS Chem Biol. 2021. PMID: 34192867 Free PMC article.
References
-
- Brosius J, Chen R. The primary structure of protein L16 located at the peptidyltransferase center of Escherichia coli ribosomes. FEBS letters. 1976;68:105–109. - PubMed
-
- Wittmann-Liebold B, Pannenbecker R. Primary structure of protein L33 from the large subunit of the Escherichia coli ribosome. FEBS letters. 1976;68:115–118. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

