Fully automatic evaluation of the corneal endothelium from in vivo confocal microscopy
- PMID: 25928199
- PMCID: PMC4451752
- DOI: 10.1186/s12880-015-0054-3
Fully automatic evaluation of the corneal endothelium from in vivo confocal microscopy
Abstract
Background: Manual and semi-automatic analyses of images, acquired in vivo by confocal microscopy, are often used to determine the quality of corneal endothelium in the human eye. These procedures are highly time consuming. Here, we present two fully automatic methods to analyze and quantify corneal endothelium imaged by in vivo white light slit-scanning confocal microscopy.
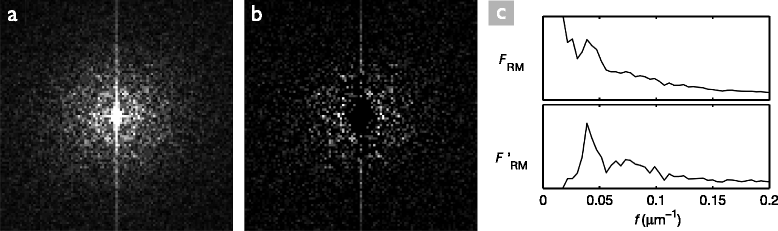
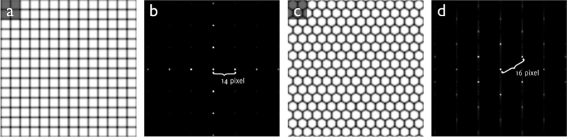
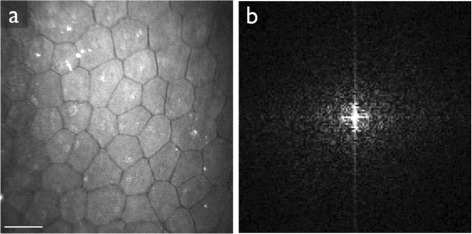
Methods: In the first approach, endothelial cell density is estimated with the help of spatial frequency analysis. We evaluate published methods, and propose a new, parameter-free method. In the second approach, based on the stochastic watershed, cells are automatically segmented and the result is used to estimate cell density, polymegathism (cell size variability) and pleomorphism (cell shape variation). We show how to determine optimal values for the three parameters of this algorithm, and compare its results to a semi-automatic delineation by a trained observer.
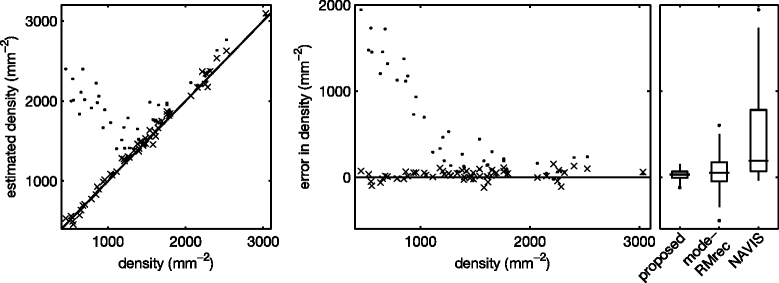
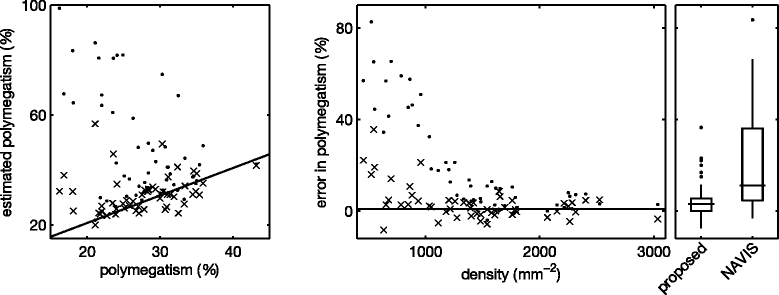
Results: The frequency analysis method proposed here is more precise than any published method. The segmentation method outperforms the fully automatic method in the NAVIS software (Nidek Technologies Srl, Padova, Italy), which significantly overestimates the number of cells for cell densities below approximately 1200 mm(-2), as well as previously published methods.
Conclusions: The methods presented here provide a significant improvement over the state of the art, and make in vivo, automated assessment of corneal endothelium more accessible. The segmentation method proposed paves the way to many possible new morphometric parameters, which can quickly and precisely be determined from the segmented image.
Figures







References
-
- Bourne WM, Nelson LR, Hodge DO. Central corneal endothelial cell changes over a ten-year period. Invest Ophthalmol Vis Sci. 1997;38(3):779–82. - PubMed
-
- Yee RW, Geroski DH, Matsuda M, Champeau EJ, Meyer LA, Edelhauser HF. Correlation of corneal endothelial pump site density, barrier function, and morphology in wound repair. Invest Ophthalmol Vis Sci. 1985;26(9):1191–2101. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

