Integrin beta-8 (ITGB8) silencing reverses gefitinib resistance of human hepatic cancer HepG2/G cell line
- PMID: 25932283
- PMCID: PMC4402930
Integrin beta-8 (ITGB8) silencing reverses gefitinib resistance of human hepatic cancer HepG2/G cell line
Abstract
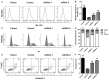
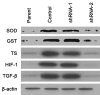
Hepatic cancer is a class of cancer that is relatively insensitive to chemotherapy, and cancers that harbor EGFR active mutations are more sensitive to EGFR-TK inhibitor such as gefitinib, which becomes the first-line treatment of this subtype of cancer. However, almost all patients treated with gefitinib will develop drug resistance. Here we show that a protein called integrin beta-8 (ITGB8) when over-expressed, is correlated with the gefitinib resistance of hepatic cancer cell line HepG2/G. After ITGB8 silencing, the drug resistance is reversed as the cell proliferation decreases and apoptosis rate increases significantly by gefitinib treatment when compared to HepG2/G. We demonstrated that multi-drug resistant proteins ABCB1, ABCC2 and ABCG2, anti-apoptosis proteins like survivin and Bcl-2, and cycle promoting protein CDK1 are involved in drug resistance of HepG2/G. Other drug-resistance relative proteins like SOD, GST, TS and HIF-1 are also modulated by ITGB8 silencing, but their role in this gefitinib resistance might be indirect. TGF beta pathway could be a critical pathway by which ITGB8 modulates the sensitivity of HepG2/G to gefitinib.
Keywords: Hepatic cancer; gefitinib resistance; integrin beta-8.
Figures




Similar articles
-
PLAUR Confers Resistance to Gefitinib Through EGFR/P-AKT/Survivin Signaling Pathway.Cell Physiol Biochem. 2018;47(5):1909-1924. doi: 10.1159/000491071. Epub 2018 Jun 29. Cell Physiol Biochem. 2018. PMID: 29961070
-
Epigenetic silencing of miR-483-3p promotes acquired gefitinib resistance and EMT in EGFR-mutant NSCLC by targeting integrin β3.Oncogene. 2018 Aug;37(31):4300-4312. doi: 10.1038/s41388-018-0276-2. Epub 2018 May 2. Oncogene. 2018. PMID: 29717264 Free PMC article.
-
Gefitinib inhibition of drug resistance to doxorubicin by inactivating ABCG2 in thyroid cancer cell lines.Arch Otolaryngol Head Neck Surg. 2007 Oct;133(10):1022-7. doi: 10.1001/archotol.133.10.1022. Arch Otolaryngol Head Neck Surg. 2007. PMID: 17938326
-
An update of the mechanisms of resistance to EGFR-tyrosine kinase inhibitors in breast cancer: Gefitinib (Iressa) -induced changes in the expression and nucleo-cytoplasmic trafficking of HER-ligands (Review).Int J Mol Med. 2007 Jul;20(1):3-10. Int J Mol Med. 2007. PMID: 17549382 Review.
-
Gefitinib: current status in the treatment of non-small cell lung cancer.Drugs Today (Barc). 2004 Oct;40(10):809-27. doi: 10.1358/dot.2004.40.10.863742. Drugs Today (Barc). 2004. PMID: 15605116 Review.
Cited by
-
Cancer-Associated Membrane Protein as Targeted Therapy for Bladder Cancer.Pharmaceutics. 2022 Oct 18;14(10):2218. doi: 10.3390/pharmaceutics14102218. Pharmaceutics. 2022. PMID: 36297654 Free PMC article. Review.
-
Predicting the clinical outcome of oral potentially malignant disorders using transcriptomic-based molecular pathology.Br J Cancer. 2021 Aug;125(3):413-421. doi: 10.1038/s41416-021-01411-z. Epub 2021 May 10. Br J Cancer. 2021. PMID: 33972745 Free PMC article.
-
Comprehensive characterization of a drug-resistance-related ceRNA network across 15 anti-cancer drug categories.Mol Ther Nucleic Acids. 2021 Feb 15;24:11-24. doi: 10.1016/j.omtn.2021.02.011. eCollection 2021 Jun 4. Mol Ther Nucleic Acids. 2021. PMID: 33738135 Free PMC article.
-
LINC01798/miR-17-5p axis regulates ITGA8 and causes changes in tumor microenvironment and stemness in lung adenocarcinoma.Front Immunol. 2023 Feb 23;14:1096818. doi: 10.3389/fimmu.2023.1096818. eCollection 2023. Front Immunol. 2023. PMID: 36911684 Free PMC article.
-
LncRNA TCF7 Promotes Epithelial Ovarian Cancer Viability, Mobility and Stemness via Regulating ITGB8.Front Oncol. 2021 Nov 17;11:649655. doi: 10.3389/fonc.2021.649655. eCollection 2021. Front Oncol. 2021. Retraction in: Front Oncol. 2023 Oct 19;13:1307324. doi: 10.3389/fonc.2023.1307324. PMID: 34868900 Free PMC article. Retracted.
References
-
- Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127:2893–2917. - PubMed
-
- Zhu Y, Zhu L, Lu L, Zhang L, Zhang G, Wang Q, Yang P. Role and mechanism of the alkylglycerone phosphate synthase in suppressing the invasion potential of human glioma and hepatic carcinoma cells in vitro. Oncol Rep. 2014;32:431–436. - PubMed
-
- Chen MJ, Zhong W, Zhang L, Zhao J, Li LY, Wang MZ. Recurrence patterns of advanced non-small cell lung cancer treated with gefitinib. Chin Med. 2013;126:2235–2241. - PubMed
-
- Karroum O, Kengen J, Gregoire V, Gallez B, Jordan BF. Tumor reoxygenation following administration of the EGFR inhibitor, gefitinib, in experimental tumors. Adv Exp Med Biol. 2013;789:265–271. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
