Calcium sensors of ciliary outer arm dynein: functions and phylogenetic considerations for eukaryotic evolution
- PMID: 25932323
- PMCID: PMC4415241
- DOI: 10.1186/s13630-015-0015-z
Calcium sensors of ciliary outer arm dynein: functions and phylogenetic considerations for eukaryotic evolution
Abstract
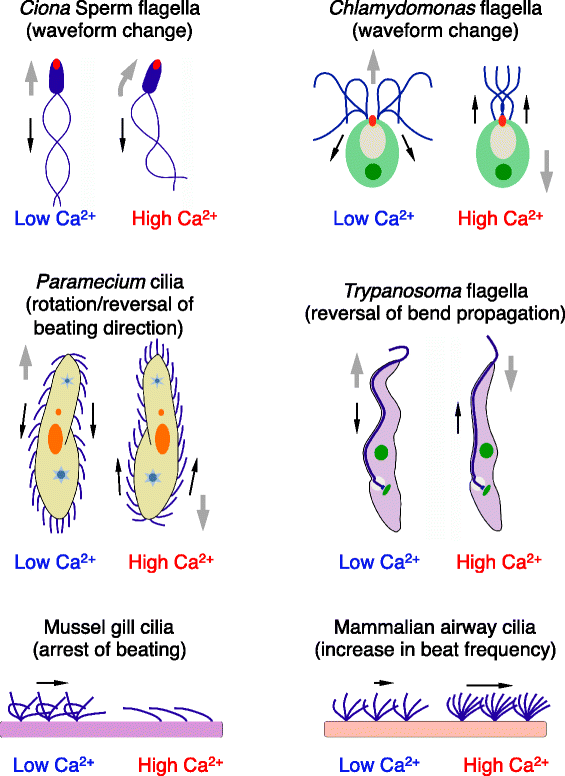
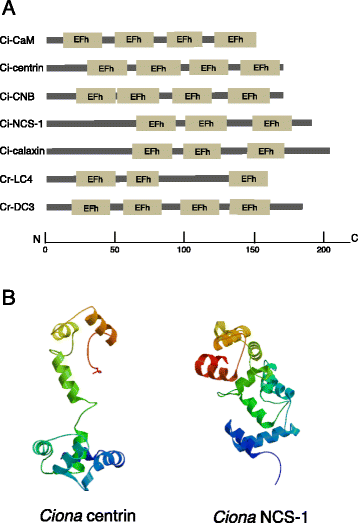
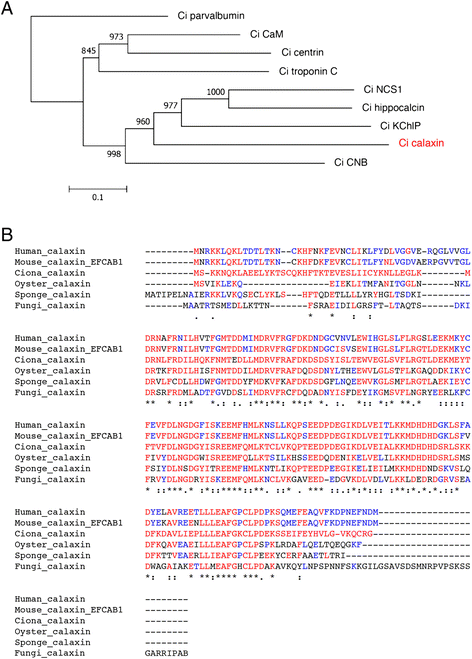
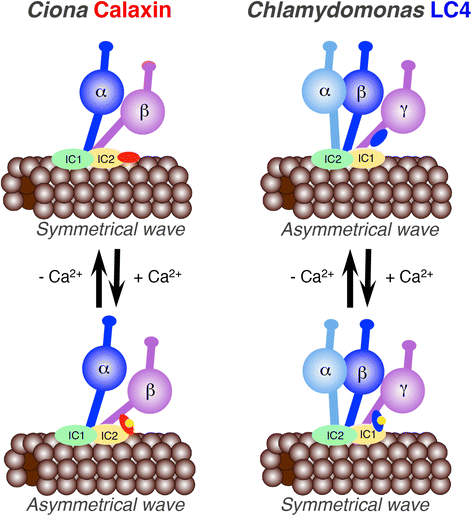
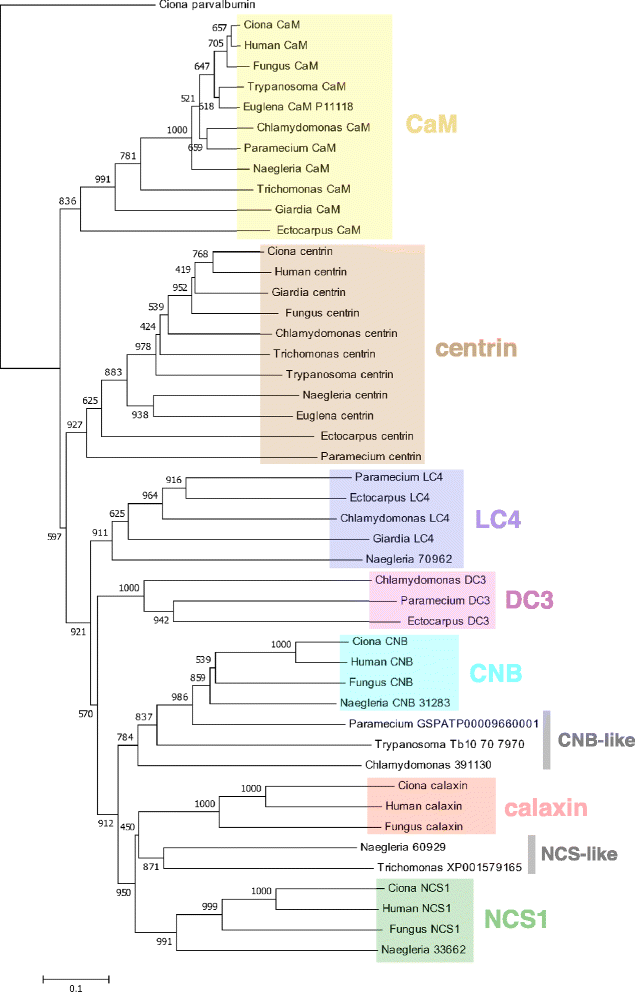
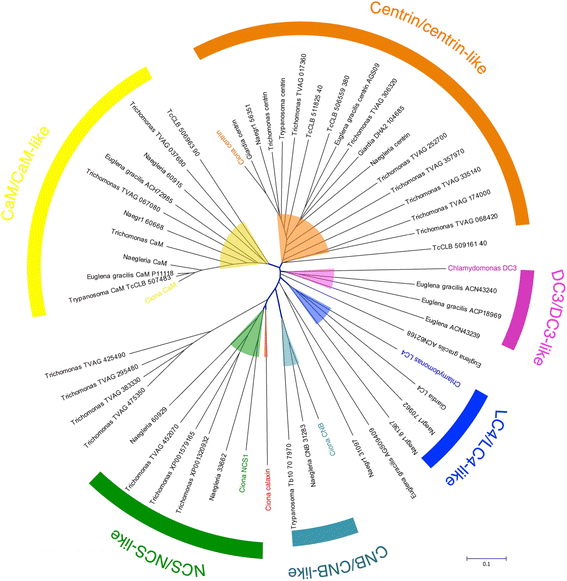
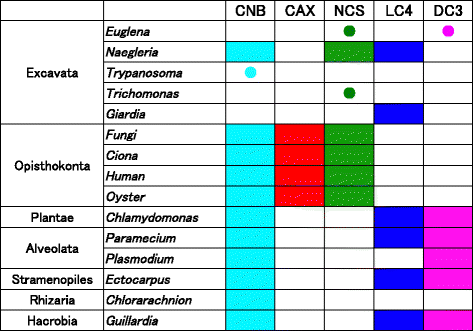
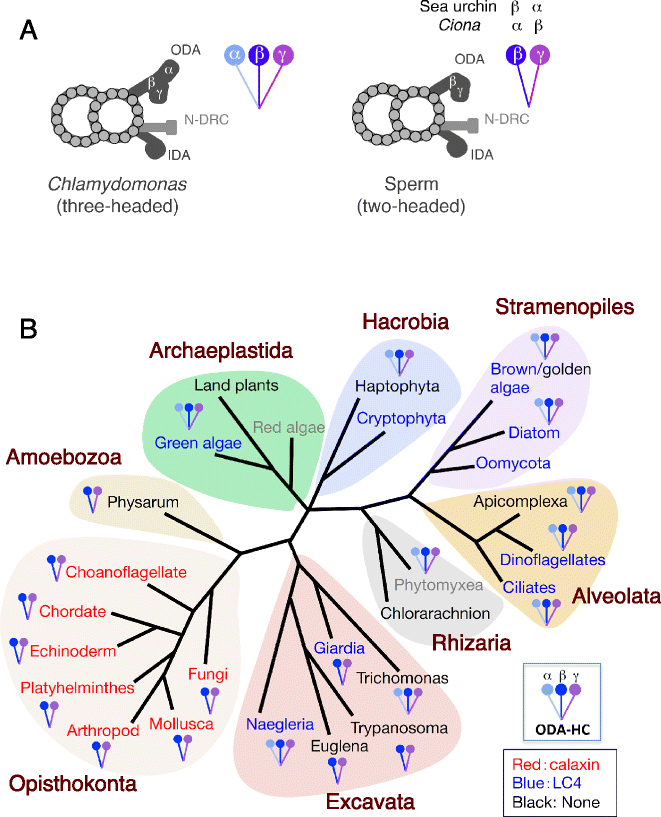
The motility of eukaryotic cilia and flagella is modulated in response to several extracellular stimuli. Ca(2+) is the most critical intracellular factor for these changes in motility, directly acting on the axonemes and altering flagellar asymmetry. Calaxin is an opisthokont-specific neuronal calcium sensor protein first described in the sperm of the ascidian Ciona intestinalis. It binds to a heavy chain of two-headed outer arm dynein in a Ca(2+)-dependent manner and regulates 'asymmetric' wave propagation at high concentrations of Ca(2+). A Ca(2+)-binding subunit of outer arm dynein in Chlamydomonas reinhardtii, the light chain 4 (LC4), which is a Ca(2+)-sensor phylogenetically different from calaxin, shows Ca(2+)-dependent binding to a heavy chain of three-headed outer arm dynein. However, LC4 appears to participate in 'symmetric' wave propagation at high concentrations of Ca(2+). LC4-type dynein light chain is present in bikonts, except for some subclasses of the Excavata. Thus, flagellar asymmetry-symmetry conversion in response to Ca(2+) concentration represents a 'mirror image' relationship between Ciona and Chlamydomonas. Phylogenetic analyses indicate the duplication, divergence, and loss of heavy chain and Ca(2+)-sensors of outer arm dynein among excavate species. These features imply a divergence point with respect to Ca(2+)-dependent regulation of outer arm dynein in cilia and flagella during the evolution of eukaryotic supergroups.
Keywords: Algae; Bikont; Calaxin; Eukaryote; Evolution; Excavate; Excavates; Fertilization; Neuronal calcium sensor; Opisthokont; Sperm.
Figures
Similar articles
-
A novel neuronal calcium sensor family protein, calaxin, is a potential Ca(2+)-dependent regulator for the outer arm dynein of metazoan cilia and flagella.Biol Cell. 2009 Feb;101(2):91-103. doi: 10.1042/BC20080032. Biol Cell. 2009. PMID: 18620543
-
Inverse relationship of Ca2+-dependent flagellar response between animal sperm and prasinophyte algae.J Plant Res. 2017 May;130(3):465-473. doi: 10.1007/s10265-017-0931-7. Epub 2017 Apr 19. J Plant Res. 2017. PMID: 28424932
-
Calaxin is required for asymmetric bend initiation and propagation in sperm flagella.Front Cell Dev Biol. 2023 Mar 16;11:1136404. doi: 10.3389/fcell.2023.1136404. eCollection 2023. Front Cell Dev Biol. 2023. PMID: 37009483 Free PMC article.
-
Composition and function of ciliary inner-dynein-arm subunits studied in Chlamydomonas reinhardtii.Cytoskeleton (Hoboken). 2021 Mar;78(3):77-96. doi: 10.1002/cm.21662. Epub 2021 Apr 28. Cytoskeleton (Hoboken). 2021. PMID: 33876572 Free PMC article. Review.
-
Functional diversity of axonemal dyneins as assessed by in vitro and in vivo motility assays of Chlamydomonas mutants.Zoolog Sci. 2014 Oct;31(10):633-44. doi: 10.2108/zs140066. Zoolog Sci. 2014. PMID: 25284382 Review.
Cited by
-
From damage response to action potentials: early evolution of neural and contractile modules in stem eukaryotes.Philos Trans R Soc Lond B Biol Sci. 2016 Jan 5;371(1685):20150043. doi: 10.1098/rstb.2015.0043. Philos Trans R Soc Lond B Biol Sci. 2016. PMID: 26598726 Free PMC article. Review.
-
The steering gaits of sperm.Philos Trans R Soc Lond B Biol Sci. 2020 Feb 17;375(1792):20190149. doi: 10.1098/rstb.2019.0149. Epub 2019 Dec 30. Philos Trans R Soc Lond B Biol Sci. 2020. PMID: 31884910 Free PMC article. Review.
-
Expression of T-Type Voltage-Gated Calcium Channel in the Cilia of Human Nasal Epithelial Cells.Int Arch Allergy Immunol. 2022;183(6):579-590. doi: 10.1159/000521765. Epub 2022 Jan 31. Int Arch Allergy Immunol. 2022. PMID: 35100604 Free PMC article.
-
Sperm dysfunction and ciliopathy.Reprod Med Biol. 2015 Oct 14;15(2):77-94. doi: 10.1007/s12522-015-0225-5. eCollection 2016 Apr. Reprod Med Biol. 2015. PMID: 29259424 Free PMC article. Review.
-
The MpCAFA gene encodes a ciliary protein required for spermatozoid motility in the liverwort Marchantia polymorpha.J Plant Res. 2025 Aug 21. doi: 10.1007/s10265-025-01666-0. Online ahead of print. J Plant Res. 2025. PMID: 40841643
References
-
- Gibbons BH, Gibbons IR. The effect of partial extraction of dynein arms on the movement of reactivated sea-urchin sperm. J Cell Sci. 1973;13:337–357. - PubMed
-
- Brokaw CL, Kamiya R. Bending patterns of Chlamydomonas flagella: IV. Mutants with defects in inner and outer dynein arms indicate differences in dynein arm function. Cell Motil Cytoskeleton. 1987;8:68–75. - PubMed
-
- Lindemann CB. Testing the geometric clutch hypothesis. Biol Cell. 2004;96:681–690. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

