Comparative genomics of Pseudomonas syringae pv. syringae strains B301D and HS191 and insights into intrapathovar traits associated with plant pathogenesis
- PMID: 25940918
- PMCID: PMC4554452
- DOI: 10.1002/mbo3.261
Comparative genomics of Pseudomonas syringae pv. syringae strains B301D and HS191 and insights into intrapathovar traits associated with plant pathogenesis
Abstract
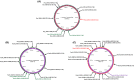
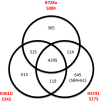
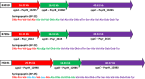
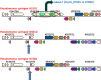
Pseudomonas syringae pv. syringae is a common plant-associated bacterium that causes diseases of both monocot and dicot plants worldwide. To help delineate traits critical to adaptation and survival in the plant environment, we generated complete genome sequences of P. syringae pv. syringae strains B301D and HS191, which represent dicot and monocot strains with distinct host specificities. Intrapathovar comparisons of the B301D (6.09 Mb) and HS191 (5.95 Mb plus a 52 kb pCG131 plasmid) genomes to the previously sequenced B728a genome demonstrated that the shared genes encompass about 83% of each genome, and include genes for siderophore biosynthesis, osmotolerance, and extracellular polysaccharide production. Between 7% and 12% of the genes are unique among the genomes, and most of the unique gene regions carry transposons, phage elements, or IS elements associated with horizontal gene transfer. Differences are observed in the type III effector composition for the three strains that likely influences host range. The HS191 genome had the largest number at 25 of effector genes, and seven effector genes are specific to this monocot strain. Toxin production is another major trait associated with virulence of P. syringae pv. syringae, and HS191 is distinguished by genes for production of syringopeptin SP25 and mangotoxin.
Keywords: Comparative genomics; Pseudomonas syringae; effector gene; genome sequence; plant pathogen.
© 2015 The Authors. MicrobiologyOpen published by John Wiley & Sons Ltd.
Figures



References
-
- Alfano JR, Charkowski AO, Deng WL, Badel JL, Petnicki-Ocwieja T, van Dijk K, et al. The Pseudomonas syringae Hrp pathogenicity island has a tripartite mosaic structure composed of a cluster of type III secretion genes bounded by exchangeable effector and conserved effector loci that contribute to parasitic fitness and pathogenicity in plants. Proc. Natl. Acad. Sci. USA. 2000;97:4856–4861. - PMC - PubMed
-
- Amrein H, Makart S, Granado J, Shakya R, Schneider-Pokorny J. Dudler R. Functional analysis of genes involved in the synthesis of syringolin A by Pseudomonas syringae pv. syringae B301 D-R. Mol. Plant Microbe Interact. 2004;17:90–97. - PubMed
-
- Ballio A, Barra D, Bossa F, Collina A, Grgurina I, Marino G, et al. Syringopeptins, new phytotoxic lipodepsipeptides of Pseudomonas syringae pv. syringae. FEBS Lett. 1991;291:109–112. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

