An alternative pluripotent state confers interspecies chimaeric competency
- PMID: 25945737
- PMCID: PMC5278765
- DOI: 10.1038/nature14413
An alternative pluripotent state confers interspecies chimaeric competency
Abstract
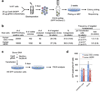
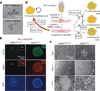
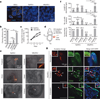
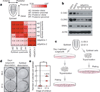
Pluripotency, the ability to generate any cell type of the body, is an evanescent attribute of embryonic cells. Transitory pluripotent cells can be captured at different time points during embryogenesis and maintained as embryonic stem cells or epiblast stem cells in culture. Since ontogenesis is a dynamic process in both space and time, it seems counterintuitive that these two temporal states represent the full spectrum of organismal pluripotency. Here we show that by modulating culture parameters, a stem-cell type with unique spatial characteristics and distinct molecular and functional features, designated as region-selective pluripotent stem cells (rsPSCs), can be efficiently obtained from mouse embryos and primate pluripotent stem cells, including humans. The ease of culturing and editing the genome of human rsPSCs offers advantages for regenerative medicine applications. The unique ability of human rsPSCs to generate post-implantation interspecies chimaeric embryos may facilitate our understanding of early human development and evolution.
Conflict of interest statement
The authors declare no competing financial interests.
Figures











Comment in
-
Stem cells: Equilibrium established.Nature. 2015 May 21;521(7552):299-300. doi: 10.1038/521299a. Nature. 2015. PMID: 25993958 No abstract available.
-
rsPSCs: A new type of pluripotent stem cells.Cell Res. 2015 Aug;25(8):889-90. doi: 10.1038/cr.2015.73. Epub 2015 Jun 16. Cell Res. 2015. PMID: 26077383 Free PMC article.
References
-
- Evans MJ, Kaufman MH. Establishment in culture of pluripotential cells from mouse embryos. Nature. 1981;292:154–156. - PubMed
-
- Martello G, Smith A. The nature of embryonic stem cells. Annu. Rev. Cell Dev. Biol. 2014;30:647–675. - PubMed
-
- Tesar PJ, et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature. 2007;448:196–199. - PubMed
-
- Brons IGM, et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature. 2007;448:191–195. - PubMed
Publication types
MeSH terms
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

