Identification and characterization of the specific murine NK cell subset supporting graft- versus-leukemia- and reducing graft- versus-host-effects
- PMID: 25949862
- PMCID: PMC4368119
- DOI: 10.4161/2162402X.2014.981483
Identification and characterization of the specific murine NK cell subset supporting graft- versus-leukemia- and reducing graft- versus-host-effects
Abstract
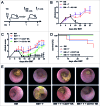
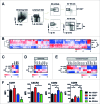
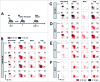
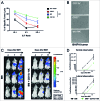
Clinical studies investigating the impact of natural killer (NK) cells in allogeneic hematopoietic stem cell transplantation settings have yielded promising results. However, NK cells are a functionally and phenotypically heterogeneous population. Therefore, we addressed the functional relevance of specific NK cell subsets distinguished by expression of CD117, CD27 and CD11b surface markers in graft-versus-leukemia (GVL)-reaction and graft-versus-host-disease (GVHD). Our results clearly demonstrate that the subset of c-Kit-CD27-CD11b+ NK cells expressed multiple cytotoxic pathway genes and provided optimal graft-versus-leukemia-effects, while significantly reducing T cell proliferation induced by allogeneic dendritic cells. Furthermore, these NK cells migrated to inflamed intestinal tissues where graft-versus-host-colitis was efficiently mitigated. For the first time, we identified the c-Kit-CD27-CD11b+ NK cell population as the specific effector NK cell subset capable of significantly diminishing GVHD in fully mismatched bone marrow transplantation settings. In conclusion, the subset of c-Kit-CD27-CD11b+ NK cells not only supports GVL, but also plays a unique role in the protection against GVHD by migrating to the peripheral GVHD target organs where they exert efficient immunoregulatory activities. These new insights demonstrate the importance of selecting the optimal NK cell subset for cellular immunotherapy following allogeneic hematopoietic stem cell transplantation.
Keywords: BMT, bone marrow transplantation; CD11b+ NK = c-Kit−CD27−CD11b+ NK cells; CD27+ NK = c-Kit−CD27+CD11b− NK cells; DP = c-Kit−CD27+CD11b+ NK cells; GVHD; GVHD, graft-versus-host disease; GVL; GVL, graft-versus-leukemia; HSCT, hematopoietic stem cell transplantation; KIR, killer cell immunoglobulin-like receptor; MLR, mixed lymphocyte reaction; NK cells; NK, natural killer; TBI, total body irradiation; c-Kit+ NK = c-Kit+CD27+CD11b− NK cells; stem cell transplantation; tumor immunology.
Figures


References
-
- Ruggeri L, Capanni M, Urbani E, Perruccio K, Shlomchik WD, Tosti A, et al. Effectiveness of donor natural killer cell alloreactivity in mismatched hematopoietic transplants. Science 2002; 295:2097-100. - PubMed
-
- Giebel S, Locatelli F, Lamparelli T, Velardi A, Davies S, Frumento G, et al. Survival advantage with KIR ligand incompatibility in hematopoietic stem cell transplantation from unrelated donors. Blood 2003; 102:814-9. - PubMed
-
- Miller JS, Soignier Y, Panoskaltsis-Mortari A, McNearney SA, Yun GH, Fautsch SK, et al. Successful adoptive transfer and in vivo expansion of human haploidentical NK cells in patients with cancer. Blood 2005; 105:3051-7. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
