Dosage compensation can buffer copy-number variation in wild yeast
- PMID: 25955966
- PMCID: PMC4448642
- DOI: 10.7554/eLife.05462
Dosage compensation can buffer copy-number variation in wild yeast
Erratum in
-
Correction: Dosage compensation can buffer copy-number variation in wild yeast.Elife. 2016 Mar 23;5:e15743. doi: 10.7554/eLife.15743. Elife. 2016. PMID: 27006204 Free PMC article. No abstract available.
Abstract
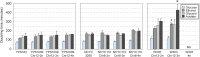
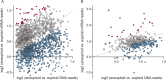
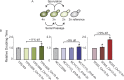
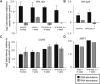
Aneuploidy is linked to myriad diseases but also facilitates organismal evolution. It remains unclear how cells overcome the deleterious effects of aneuploidy until new phenotypes evolve. Although laboratory strains are extremely sensitive to aneuploidy, we show here that aneuploidy is common in wild yeast isolates, which show lower-than-expected expression at many amplified genes. We generated diploid strain panels in which cells carried two, three, or four copies of the affected chromosomes, to show that gene-dosage compensation functions at >30% of amplified genes. Genes subject to dosage compensation are under higher expression constraint in wild populations-but they show elevated rates of gene amplification, suggesting that copy-number variation is buffered at these genes. We find that aneuploidy provides a clear ecological advantage to oak strain YPS1009, by amplifying a causal gene that escapes dosage compensation. Our work presents a model in which dosage compensation buffers gene amplification through aneuploidy to provide a natural, but likely transient, route to rapid phenotypic evolution.
Keywords: S. cerevisiae; aneuploidy; computational biology; copy number variation; evolutionary biology; gene expression; genomics; natural variation; stress resistance; systems biology.
Conflict of interest statement
The authors declare that no competing interests exist.
Figures



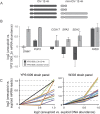


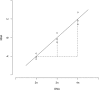
Comment in
-
Aneuploidy: Tolerating Tolerance.Curr Biol. 2015 Aug 31;25(17):R771-3. doi: 10.1016/j.cub.2015.06.056. Curr Biol. 2015. PMID: 26325142
References
-
- Benjamini Y, Hochberg Y. Controllign the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society. 1995;57:289–300.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

