Mechanisms of cancer cell killing by the adenovirus E4orf4 protein
- PMID: 25961489
- PMCID: PMC4452909
- DOI: 10.3390/v7052334
Mechanisms of cancer cell killing by the adenovirus E4orf4 protein
Abstract
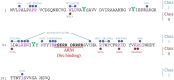
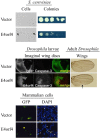
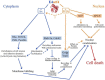
During adenovirus (Ad) replication the Ad E4orf4 protein regulates progression from the early to the late phase of infection. However, when E4orf4 is expressed alone outside the context of the virus it induces a non-canonical mode of programmed cell death, which feeds into known cell death pathways such as apoptosis or necrosis, depending on the cell line tested. E4orf4-induced cell death has many interesting and unique features including a higher susceptibility of cancer cells to E4orf4-induced cell killing compared with normal cells, caspase-independence, a high degree of evolutionary conservation of the signaling pathways, a link to perturbations of the cell cycle, and involvement of two distinct cell death programs, in the nucleus and in the cytoplasm. Several E4orf4-interacting proteins including its major partners, protein phosphatase 2A (PP2A) and Src family kinases, contribute to induction of cell death. The various features of E4orf4-induced cell killing as well as studies to decipher the underlying mechanisms are described here. Many explanations for the cancer specificity of E4orf4-induced cell death have been proposed, but a full understanding of the reasons for the different susceptibility of cancer and normal cells to killing by E4orf4 will require a more detailed analysis of the complex E4orf4 signaling network. An improved understanding of the mechanisms involved in this unique mode of programmed cell death may aid in design of novel E4orf4-based cancer therapeutics.
Figures

Similar articles
-
The Adenovirus E4orf4 Protein Provides a Novel Mechanism for Inhibition of the DNA Damage Response.PLoS Pathog. 2016 Feb 11;12(2):e1005420. doi: 10.1371/journal.ppat.1005420. eCollection 2016 Feb. PLoS Pathog. 2016. PMID: 26867009 Free PMC article.
-
Induction of cancer-specific cell death by the adenovirus E4orf4 protein.Adv Exp Med Biol. 2014;818:61-97. doi: 10.1007/978-1-4471-6458-6_4. Adv Exp Med Biol. 2014. PMID: 25001532 Review.
-
NTPDASE4 gene products cooperate with the adenovirus E4orf4 protein through PP2A-dependent and -independent mechanisms and contribute to induction of cell death.J Virol. 2014 Jun;88(11):6318-28. doi: 10.1128/JVI.00381-14. Epub 2014 Mar 26. J Virol. 2014. PMID: 24672025 Free PMC article.
-
Distinct cell death pathways triggered by the adenovirus early region 4 ORF 4 protein.J Cell Biol. 2002 Aug 5;158(3):519-28. doi: 10.1083/jcb.200201106. Epub 2002 Aug 5. J Cell Biol. 2002. PMID: 12163473 Free PMC article.
-
Biology of the adenovirus E4orf4 protein: from virus infection to cancer cell death.FEBS Lett. 2020 Jun;594(12):1891-1917. doi: 10.1002/1873-3468.13704. Epub 2019 Dec 12. FEBS Lett. 2020. PMID: 31792953 Review.
Cited by
-
Identification of the Potential Role of the E4orf4 Protein in Adenovirus A, B, C, and D Groups in Cancer Therapy: Computational Approaches.Mol Biotechnol. 2024 Sep 13. doi: 10.1007/s12033-024-01278-4. Online ahead of print. Mol Biotechnol. 2024. PMID: 39269574
-
Selective elimination of cancer cells by the adenovirus E4orf4 protein in a Drosophila cancer model: a new paradigm for cancer therapy.Cell Death Dis. 2019 Jun 11;10(6):455. doi: 10.1038/s41419-019-1680-4. Cell Death Dis. 2019. PMID: 31186403 Free PMC article.
-
Biphasic Functional Interaction between the Adenovirus E4orf4 Protein and DNA-PK.J Virol. 2019 May 1;93(10):e01365-18. doi: 10.1128/JVI.01365-18. Print 2019 May 15. J Virol. 2019. PMID: 30842317 Free PMC article.
-
The Adenovirus E4orf4 Protein Provides a Novel Mechanism for Inhibition of the DNA Damage Response.PLoS Pathog. 2016 Feb 11;12(2):e1005420. doi: 10.1371/journal.ppat.1005420. eCollection 2016 Feb. PLoS Pathog. 2016. PMID: 26867009 Free PMC article.
-
An Interaction with PARP-1 and Inhibition of Parylation Contribute to Attenuation of DNA Damage Signaling by the Adenovirus E4orf4 Protein.J Virol. 2019 Sep 12;93(19):e02253-18. doi: 10.1128/JVI.02253-18. Print 2019 Oct 1. J Virol. 2019. PMID: 31315986 Free PMC article.
References
-
- Mannervik M., Fan S., Strom A.C., Helin K., Akusjarvi G. Adenovirus E4 open reading frame 4-induced dephosphorylation inhibits E1A activation of the E2 promoter and E2F-1-mediated transactivation independently of the retinoblastoma tumor suppressor protein. Virology. 1999;256:313–321. doi: 10.1006/viro.1999.9663. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

