A nuclear role for the respiratory enzyme CLK-1 in regulating mitochondrial stress responses and longevity
- PMID: 25961505
- PMCID: PMC4539581
- DOI: 10.1038/ncb3170
A nuclear role for the respiratory enzyme CLK-1 in regulating mitochondrial stress responses and longevity
Abstract
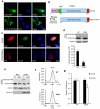
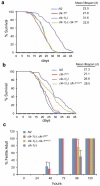
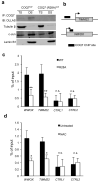
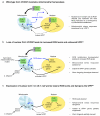
The coordinated regulation of mitochondrial and nuclear activities is essential for cellular respiration and its disruption leads to mitochondrial dysfunction, a hallmark of ageing. Mitochondria communicate with nuclei through retrograde signalling pathways that modulate nuclear gene expression to maintain mitochondrial homeostasis. The monooxygenase CLK-1 (human homologue COQ7) was previously reported to be mitochondrial, with a role in respiration and longevity. We have uncovered a distinct nuclear form of CLK-1 that independently regulates lifespan. Nuclear CLK-1 mediates a retrograde signalling pathway that is conserved from Caenorhabditis elegans to humans and is responsive to mitochondrial reactive oxygen species, thus acting as a barometer of oxidative metabolism. We show that, through modulation of gene expression, the pathway regulates both mitochondrial reactive oxygen species metabolism and the mitochondrial unfolded protein response. Our results demonstrate that a respiratory enzyme acts in the nucleus to control mitochondrial stress responses and longevity.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

