Dot1 histone methyltransferases share a distributive mechanism but have highly diverged catalytic properties
- PMID: 25965993
- PMCID: PMC4650758
- DOI: 10.1038/srep09824
Dot1 histone methyltransferases share a distributive mechanism but have highly diverged catalytic properties
Abstract
The conserved histone methyltransferase Dot1 establishes an H3K79 methylation pattern consisting of mono-, di- and trimethylation states on histone H3 via a distributive mechanism. This mechanism has been shown to be important for the regulation of the different H3K79 methylation states in yeast. Dot1 enzymes in yeast, Trypanosoma brucei (TbDot1A and TbDot1B, which methylate H3K76) and human (hDot1L) generate very divergent methylation patterns. To understand how these species-specific methylation patterns are generated, the methylation output of the Dot1 enzymes was compared by expressing them in yeast at various expression levels. Computational simulations based on these data showed that the Dot1 enzymes have highly distinct catalytic properties, but share a distributive mechanism. The mechanism of methylation and the distinct rate constants have implications for the regulation of H3K79/K76 methylation. A mathematical model of H3K76 methylation during the trypanosome cell cycle suggests that temporally-regulated consecutive action of TbDot1A and TbDot1B is required for the observed regulation of H3K76 methylation states.
Figures

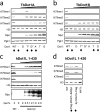
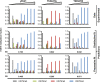
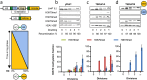
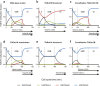
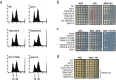
References
-
- Van Leeuwen F., Gafken P. R. & Gottschling D. E. Dot1p modulates silencing in yeast by methylation of the nucleosome core. Cell 109, 745–56 (2002). - PubMed
-
- Rusche L. N., Kirchmaier A. L. & Rine J. The establishment, inheritance, and function of silenced chromatin in Saccharomyces cerevisiae. Annu. Rev. Biochem. 72, 481–516 (2003). - PubMed
-
- McLean C. M., Karemaker I. D. & van Leeuwen F. The emerging roles of DOT1L in leukemia and normal development. Leukemia 28, 2131–8 (2014). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

