Visualizing translocation dynamics and nascent transcript errors in paused RNA polymerases in vivo
- PMID: 25976475
- PMCID: PMC4457086
- DOI: 10.1186/s13059-015-0666-5
Visualizing translocation dynamics and nascent transcript errors in paused RNA polymerases in vivo
Erratum in
-
Erratum to: Visualizing translocation dynamics and nascent transcript errors in paused RNA polymerases in vivo.Genome Biol. 2015 Dec 2;16:270. doi: 10.1186/s13059-015-0845-4. Genome Biol. 2015. PMID: 26631078 Free PMC article. No abstract available.
Abstract
Background: Transcription elongation is frequently interrupted by pausing signals in DNA, with downstream effects on gene expression. Transcription errors also induce prolonged pausing, which can lead to a destabilized genome by interfering with DNA replication. Mechanisms of pausing associated with translocation blocks and misincorporation have been characterized in vitro, but not in vivo.
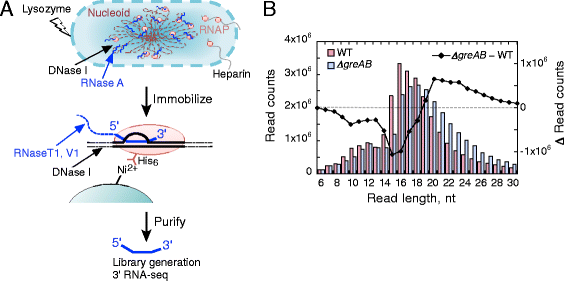
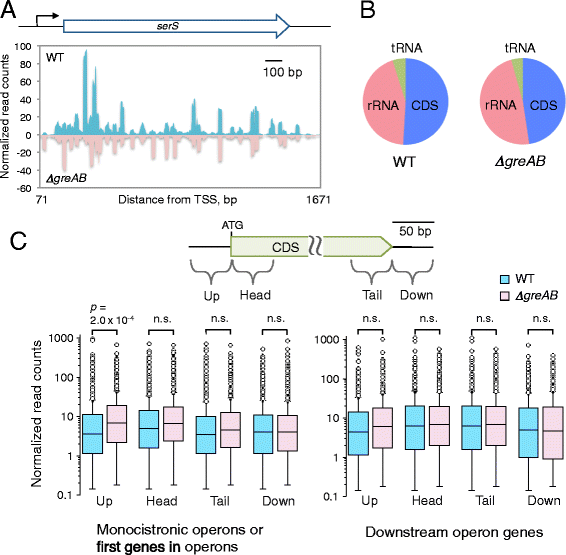
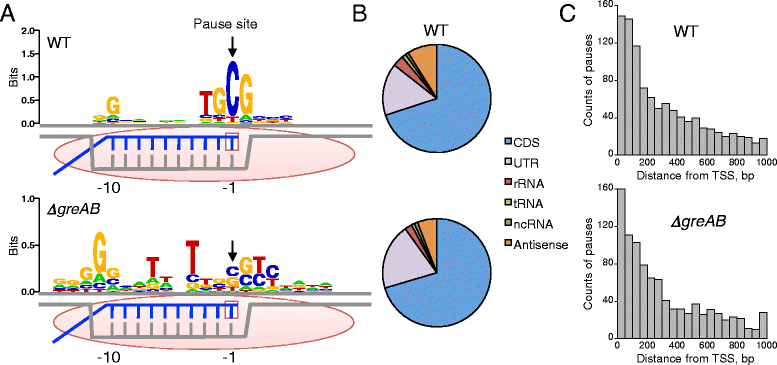
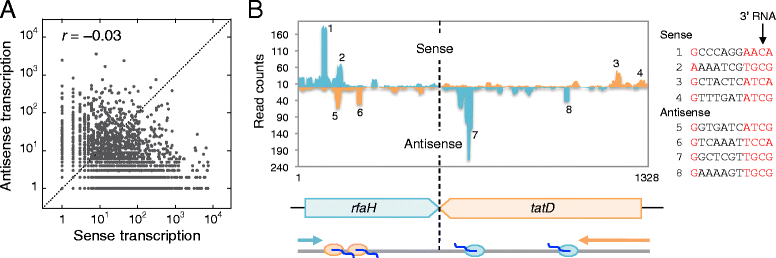
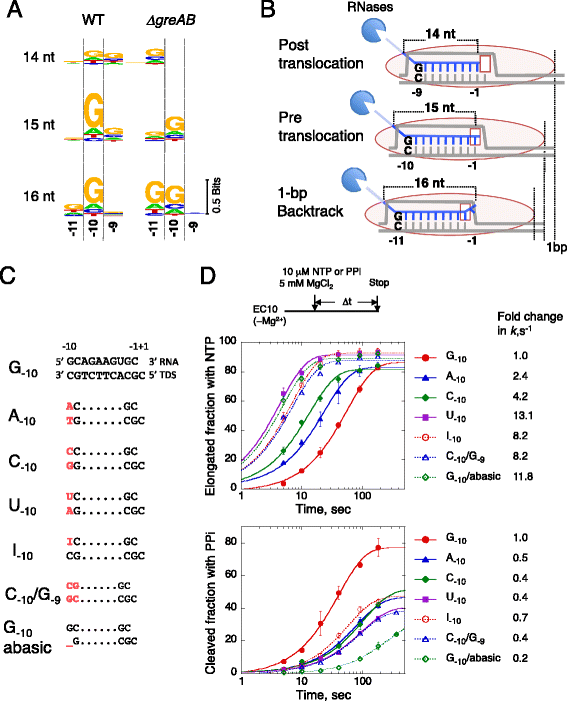
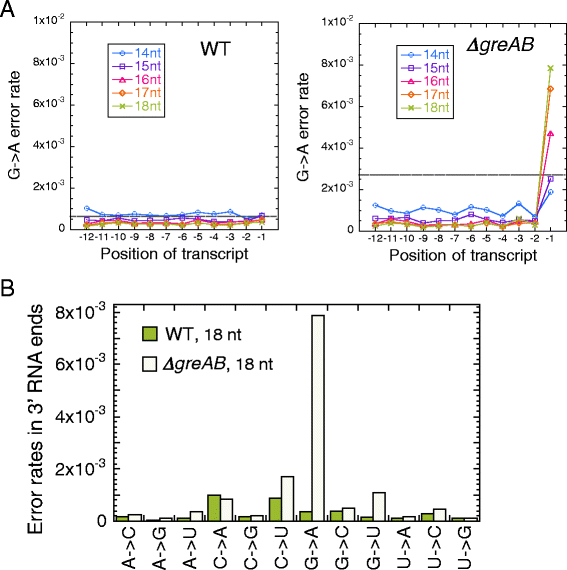
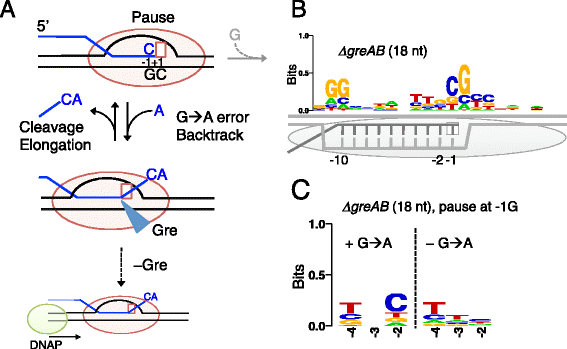
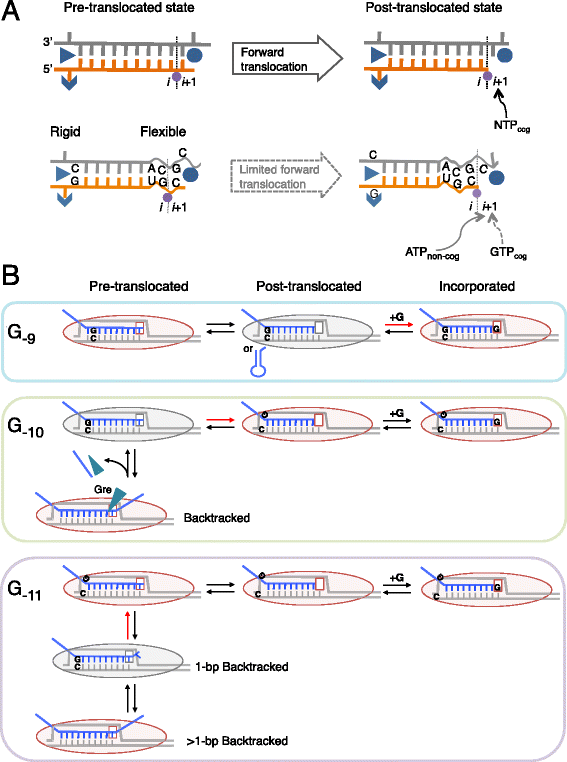
Results: We investigate the pausing pattern of RNA polymerase (RNAP) in Escherichia coli by a novel approach, combining native elongating transcript sequencing (NET-seq) with RNase footprinting of the transcripts (RNET-seq). We reveal that the G-dC base pair at the 5' end of the RNA-DNA hybrid interferes with RNAP translocation. The distance between the 5' G-dC base pair and the 3' end of RNA fluctuates over a three-nucleotide width. Thus, the G-dC base pair can induce pausing in post-translocated, pre-translocated, and backtracked states of RNAP. Additionally, a CpG sequence of the template DNA strand spanning the active site of RNAP inhibits elongation and induces G-to-A errors, which leads to backtracking of RNAP. Gre factors efficiently proofread the errors and rescue the backtracked complexes. We also find that pausing events are enriched in the 5' untranslated region and antisense transcription of mRNA genes and are reduced in rRNA genes.
Conclusions: In E. coli, robust transcriptional pausing involves RNAP interaction with G-dC at the upstream end of the RNA-DNA hybrid, which interferes with translocation. CpG DNA sequences induce transcriptional pausing and G-to-A errors.
Figures
References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

