Real-time monitoring of cell protrusion dynamics by impedance responses
- PMID: 25976978
- PMCID: PMC4432390
- DOI: 10.1038/srep10206
Real-time monitoring of cell protrusion dynamics by impedance responses
Abstract
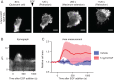
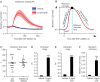
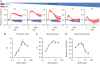
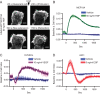
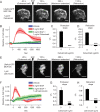
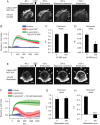
Cellular protrusions are highly dynamic structures involved in fundamental processes, including cell migration and invasion. For a cell to migrate, its leading edge must form protrusions, and then adhere or retract. The spatial and temporal coordination of protrusions and retraction is yet to be fully understood. The study of protrusion dynamics mainly relies on live-microscopy often coupled to fluorescent labeling. Here we report the use of an alternative, label-free, quantitative and rapid assay to analyze protrusion dynamics in a cell population based on the real-time recording of cell activity by means of electronic sensors. Cells are seeded on a plate covered with electrodes and their shape changes map into measured impedance variations. Upon growth factor stimulation the impedance increases due to protrusive activity and decreases following retraction. Compared to microscopy-based methods, impedance measurements are suitable to high-throughput studies on different cell lines, growth factors and chemical compounds. We present data indicating that this assay lends itself to dissect the biochemical signaling pathways controlling adhesive protrusions. Indeed, we show that the protrusion phase is sustained by actin polymerization, directly driven by growth factor stimulation. Contraction instead mainly relies on myosin action, pointing at a pivotal role of myosin in lamellipodia retraction.
Figures
Similar articles
-
ERK reinforces actin polymerization to power persistent edge protrusion during motility.Sci Signal. 2015 May 19;8(377):ra47. doi: 10.1126/scisignal.aaa8859. Sci Signal. 2015. PMID: 25990957 Free PMC article.
-
Cell mechanics control rapid transitions between blebs and lamellipodia during migration.Proc Natl Acad Sci U S A. 2012 Sep 4;109(36):14434-9. doi: 10.1073/pnas.1207968109. Epub 2012 Jul 11. Proc Natl Acad Sci U S A. 2012. PMID: 22786929 Free PMC article.
-
Hepatic stellate cell protrusions couple platelet-derived growth factor-BB to chemotaxis.Hepatology. 2007 Jun;45(6):1446-53. doi: 10.1002/hep.21606. Hepatology. 2007. PMID: 17465006
-
Pulling back to move forward.Cell. 2004 Feb 6;116(3):357-8. doi: 10.1016/s0092-8674(04)00123-0. Cell. 2004. PMID: 15016370 Review.
-
Life at the leading edge.Cell. 2011 Jun 24;145(7):1012-22. doi: 10.1016/j.cell.2011.06.010. Cell. 2011. PMID: 21703446 Review.
Cited by
-
Development of a test that measures real-time HER2 signaling function in live breast cancer cell lines and primary cells.BMC Cancer. 2017 Mar 16;17(1):199. doi: 10.1186/s12885-017-3181-0. BMC Cancer. 2017. PMID: 28302091 Free PMC article.
-
MRCKα is activated by caspase cleavage to assemble an apical actin ring for epithelial cell extrusion.J Cell Biol. 2018 Jan 2;217(1):231-249. doi: 10.1083/jcb.201703044. Epub 2017 Nov 21. J Cell Biol. 2018. PMID: 29162624 Free PMC article.
-
A new temperature-dependent strategy to modulate the epidermal growth factor receptor.Biomaterials. 2018 Nov;183:319-330. doi: 10.1016/j.biomaterials.2018.07.063. Epub 2018 Aug 11. Biomaterials. 2018. PMID: 30196151 Free PMC article.
-
RSU-1 Maintains Integrity of Caenorhabditis elegans Vulval Muscles by Regulating α-Actinin.G3 (Bethesda). 2020 Jul 7;10(7):2507-2517. doi: 10.1534/g3.120.401185. G3 (Bethesda). 2020. PMID: 32461202 Free PMC article.
-
PLEKHG5 regulates autophagy, survival and MGMT expression in U251-MG glioblastoma cells.Sci Rep. 2020 Dec 14;10(1):21858. doi: 10.1038/s41598-020-77958-3. Sci Rep. 2020. PMID: 33318498 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

