Recognition of DNA Termini by the C-Terminal Region of the Ku80 and the DNA-Dependent Protein Kinase Catalytic Subunit
- PMID: 25978375
- PMCID: PMC4433226
- DOI: 10.1371/journal.pone.0127321
Recognition of DNA Termini by the C-Terminal Region of the Ku80 and the DNA-Dependent Protein Kinase Catalytic Subunit
Abstract
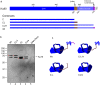
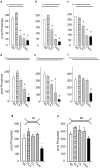
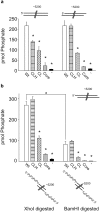
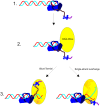
DNA double strand breaks (DSBs) can be generated by endogenous cellular processes or exogenous agents in mammalian cells. These breaks are highly variable with respect to DNA sequence and structure and all are recognized in some context by the DNA-dependent protein kinase (DNA-PK). DNA-PK is a critical component necessary for the recognition and repair of DSBs via non-homologous end joining (NHEJ). Previously studies have shown that DNA-PK responds differentially to variations in DSB structure, but how DNA-PK senses differences in DNA substrate sequence and structure is unknown. Here we explore the enzymatic mechanisms by which DNA-PK is activated by various DNA substrates and provide evidence that the DNA-PK is differentially activated by DNA structural variations as a function of the C-terminal region of Ku80. Discrimination based on terminal DNA sequence variations, on the other hand, is independent of the Ku80 C-terminal interactions and likely results exclusively from DNA-dependent protein kinase catalytic subunit interactions with the DNA. We also show that sequence differences in DNA termini can drastically influence DNA repair through altered DNA-PK activation. These results indicate that even subtle differences in DNA substrates influence DNA-PK activation and ultimately the efficiency of DSB repair.
Conflict of interest statement
Figures



References
-
- Drouet J, Frit P, Delteil C, de Villartay JP, Salles B, Calsou P (2006) Interplay between Ku, artemis, and the DNA-dependent protein kinase catalytic subunit at DNA ends. J Biol Chem 281: 27784–27793. - PubMed
-
- Furuta T, Redon C, Pilch D, Sedelnikova O, Kohlhagen G, Kirchgessner CU, et al. (2002) ATR- and DNA-PK- dependent phosphorylation of histone H2AX by replication-mediated DNA double-strand break induced by camptothecin. Gastroenterology 122: S918.
-
- Reitsema T, Klokov D, Banath JP, Olive PL (2005) DNA-PK is responsible for enhanced phosphorylation of histone H2AX under hypertonic conditions. Dna Repair 4: 1172–1181. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

