Identification of a 6-cytokine prognostic signature in patients with primary glioblastoma harboring M2 microglia/macrophage phenotype relevance
- PMID: 25978454
- PMCID: PMC4433225
- DOI: 10.1371/journal.pone.0126022
Identification of a 6-cytokine prognostic signature in patients with primary glioblastoma harboring M2 microglia/macrophage phenotype relevance
Abstract
Background: Glioblastomas (GBM) are comprised of a heterogeneous population of tumor cells, immune cells, and extracellular matrix. Interactions among these different cell types and pro-/anti-inflammatory cytokines may promote tumor development and progression.
Aims: The objective of this study was to develop a cytokine-related gene signature to improve outcome prediction for patients with primary GBM.
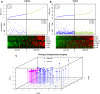
Methods: Here, we used Cox regression and risk-score analysis to develop a cytokine-related gene signature in primary GBMs from the whole transcriptome sequencing profile of the Chinese Glioma Genome Atlas (CGGA) database (n=105). We also examined differences in immune cell phenotype and immune factor expression between the high-risk and low-risk groups.
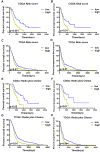
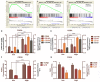
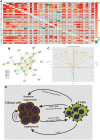
Results: Cytokine-related genes were ranked based on their ability to predict survival in the CGGA database. The six genes showing the strongest predictive value were CXCL10, IL17R, CCR2, IL17B, IL10RB, and CCL2. Patients with a high-risk score had poor overall survival and progression-free survival. Additionally, the high-risk group was characterized by increased mRNA expression of M2 microglia/macrophage markers and elevated levels of IL10 and TGFβ1.
Conclusion: The six cytokine-related gene signature is sufficient to predict survival and to identify a subgroup of primary GBM exhibiting the M2 cell phenotype.
Conflict of interest statement
Figures
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

