Robust Expansion of Human Pluripotent Stem Cells: Integration of Bioprocess Design With Transcriptomic and Metabolomic Characterization
- PMID: 25979863
- PMCID: PMC4479622
- DOI: 10.5966/sctm.2014-0270
Robust Expansion of Human Pluripotent Stem Cells: Integration of Bioprocess Design With Transcriptomic and Metabolomic Characterization
Abstract
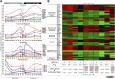
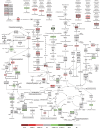
: Human embryonic stem cells (hESCs) have an enormous potential as a source for cell replacement therapies, tissue engineering, and in vitro toxicology applications. The lack of standardized and robust bioprocesses for hESC expansion has hindered the application of hESCs and their derivatives in clinical settings. We developed a robust and well-characterized bioprocess for hESC expansion under fully defined conditions and explored the potential of transcriptomic and metabolomic tools for a more comprehensive assessment of culture system impact on cell proliferation, metabolism, and phenotype. Two different hESC lines (feeder-dependent and feeder-free lines) were efficiently expanded on xeno-free microcarriers in stirred culture systems. Both hESC lines maintained the expression of stemness markers such as Oct-4, Nanog, SSEA-4, and TRA1-60 and the ability to spontaneously differentiate into the three germ layers. Whole-genome transcriptome profiling revealed a phenotypic convergence between both hESC lines along the expansion process in stirred-tank bioreactor cultures, providing strong evidence of the robustness of the cultivation process to homogenize cellular phenotype. Under low-oxygen tension, results showed metabolic rearrangement with upregulation of the glycolytic machinery favoring an anaerobic glycolysis Warburg-effect-like phenotype, with no evidence of hypoxic stress response, in contrast to two-dimensional culture. Overall, we report a standardized expansion bioprocess that can guarantee maximal product quality. Furthermore, the "omics" tools used provided relevant findings on the physiological and metabolic changes during hESC expansion in environmentally controlled stirred-tank bioreactors, which can contribute to improved scale-up production systems.
Significance: The clinical application of human pluripotent stem cells (hPSCs) has been hindered by the lack of robust protocols able to sustain production of high cell numbers, as required for regenerative medicine. In this study, a strategy was developed for the expansion of human embryonic stem cells in well-defined culture conditions using microcarrier technology and stirred-tank bioreactors. The use of transcriptomic and metabolic tools allowed detailed characterization of the cell-based product and showed a phenotypic convergence between both hESC lines along the expansion process. This study provided valuable insights into the metabolic hallmarks of hPSC expansion and new information to guide bioprocess design and media optimization for the production of cells with higher quantity and improved quality, which are requisite for translation to the clinic.
Keywords: Human pluripotent stem cells; Metabolic profiling; Stirred-tank bioreactors; Transcriptional profiling; Xeno-free microcarriers.
©AlphaMed Press.
Figures




References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

