The endogenous molecular clock orchestrates the temporal separation of substrate metabolism in skeletal muscle
- PMID: 26000164
- PMCID: PMC4440511
- DOI: 10.1186/s13395-015-0039-5
The endogenous molecular clock orchestrates the temporal separation of substrate metabolism in skeletal muscle
Abstract
Background: Skeletal muscle is a major contributor to whole-body metabolism as it serves as a depot for both glucose and amino acids, and is a highly metabolically active tissue. Within skeletal muscle exists an intrinsic molecular clock mechanism that regulates the timing of physiological processes. A key function of the clock is to regulate the timing of metabolic processes to anticipate time of day changes in environmental conditions. The purpose of this study was to identify metabolic genes that are expressed in a circadian manner and determine if these genes are regulated downstream of the intrinsic molecular clock by assaying gene expression in an inducible skeletal muscle-specific Bmal1 knockout mouse model (iMS-Bmal1 (-/-) ).
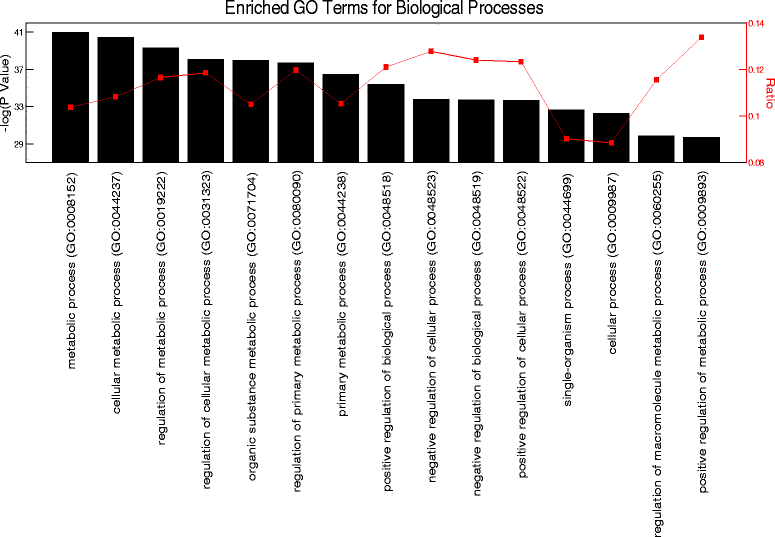
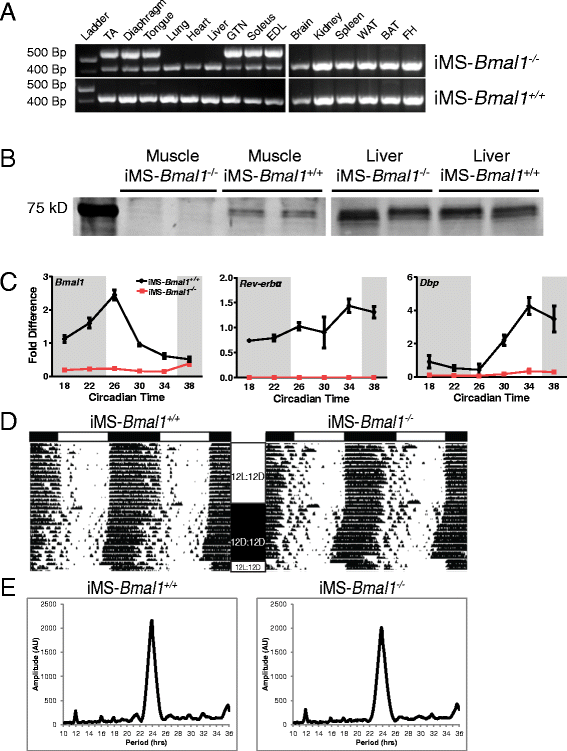
Methods: We used circadian statistics to analyze a publicly available, high-resolution time-course skeletal muscle expression dataset. Gene ontology analysis was utilized to identify enriched biological processes in the skeletal muscle circadian transcriptome. We generated a tamoxifen-inducible skeletal muscle-specific Bmal1 knockout mouse model and performed a time-course microarray experiment to identify gene expression changes downstream of the molecular clock. Wheel activity monitoring was used to assess circadian behavioral rhythms in iMS-Bmal1 (-/-) and control iMS-Bmal1 (+/+) mice.
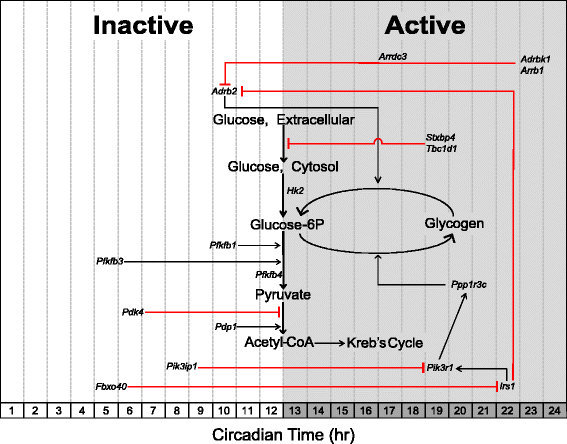
Results: The skeletal muscle circadian transcriptome was highly enriched for metabolic processes. Acrophase analysis of circadian metabolic genes revealed a temporal separation of genes involved in substrate utilization and storage over a 24-h period. A number of circadian metabolic genes were differentially expressed in the skeletal muscle of the iMS-Bmal1 (-/-) mice. The iMS-Bmal1 (-/-) mice displayed circadian behavioral rhythms indistinguishable from iMS-Bmal1 (+/+) mice. We also observed a gene signature indicative of a fast to slow fiber-type shift and a more oxidative skeletal muscle in the iMS-Bmal1 (-/-) model.
Conclusions: These data provide evidence that the intrinsic molecular clock in skeletal muscle temporally regulates genes involved in the utilization and storage of substrates independent of circadian activity. Disruption of this mechanism caused by phase shifts (that is, social jetlag) or night eating may ultimately diminish skeletal muscle's ability to efficiently maintain metabolic homeostasis over a 24-h period.
Keywords: Anabolic; Bmal1; Catabolic; Circadian; Metabolism; Molecular clock; Rev-erbα; Skeletal muscle; Temporal separation.
Figures



References
-
- Hoppeler H, Fluck M. Normal mammalian skeletal muscle and its phenotypic plasticity. J Exp Biol. 2002;205:2143–52. - PubMed
-
- Frontera WR, Ochala J. Skeletal muscle, a brief review of structure and function: Calcif Tissue Int. 2014. - PubMed
-
- Rall JA. Energetic aspects of skeletal muscle contraction: implications of fiber types. Exerc Sport Sci Rev. 1985;13:33–74. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

