Hokkaido genotype of Puumala virus in the grey red-backed vole (Myodes rufocanus) and northern red-backed vole (Myodes rutilus) in Siberia
- PMID: 26003760
- PMCID: PMC4871597
- DOI: 10.1016/j.meegid.2015.05.021
Hokkaido genotype of Puumala virus in the grey red-backed vole (Myodes rufocanus) and northern red-backed vole (Myodes rutilus) in Siberia
Abstract
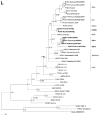
Three species of Myodes voles known to harbor hantaviruses include the bank vole (Myodes glareolus), which serves as the reservoir host of Puumala virus (PUUV), the prototype arvicolid rodent-borne hantavirus causing hemorrhagic fever with renal syndrome (HFRS) in Europe, and the grey red-backed vole (Myodes rufocanus) and royal vole (Myodes regulus) which carry two PUUV-like hantaviruses, designated Hokkaido virus (HOKV) and Muju virus (MUJV), respectively. To ascertain the hantavirus harbored by the northern red-backed vole (Myodes rutilus), we initially screened sera from 233 M. rutilus, as well as from 90 M. rufocanus and 110 M. glareolus, captured in western and eastern Siberia during June 2007 to October 2009, for anti-hantaviral antibodies. Thereafter, lung tissues from 44 seropositive voles were analyzed for hantavirus RNA by reverse transcription-polymerase chain reaction. Partial L-, M- and S-segment sequences, detected in M. rutilus and M. rufocanus, were closely related to HOKV, differing from previously published L-, M- and S-segment sequences of HOKV by 17.8-20.2%, 15.9-23.4% and 15.0-17.0% at the nucleotide level and 2.6-7.9%, 1.3-6.3% and 1.2-4.0% at the amino acid level, respectively. Alignment and comparison of hantavirus sequences from M. glareolus trapped in Tyumen Oblast showed very high sequence similarity to the Omsk lineage of PUUV. Phylogenetic analysis, using neighbor-joining, maximal likelihood and Bayesian methods, showed that HOKV strains shared a common ancestry with PUUV and exhibited geographic-specific clustering. This report provides the first molecular evidence that both M. rutilus and M. rufocanus harbor HOKV, which might represent a genetic variant of PUUV.
Keywords: Hantavirus; Myodes voles; Puumala virus; Siberia.
Copyright © 2015 Elsevier B.V. All rights reserved.
Figures



Similar articles
-
Muju virus, harbored by Myodes regulus in Korea, might represent a genetic variant of Puumala virus, the prototype arvicolid rodent-borne hantavirus.Viruses. 2014 Apr 14;6(4):1701-14. doi: 10.3390/v6041701. Viruses. 2014. PMID: 24736214 Free PMC article.
-
Isolation of Hokkaido virus, genus Hantavirus, using a newly established cell line derived from the kidney of the grey red-backed vole (Myodes rufocanus bedfordiae).J Gen Virol. 2012 Oct;93(Pt 10):2237-2246. doi: 10.1099/vir.0.045377-0. Epub 2012 Jul 12. J Gen Virol. 2012. PMID: 22791608
-
A highly divergent Puumala virus lineage in southern Poland.Arch Virol. 2017 May;162(5):1177-1185. doi: 10.1007/s00705-016-3200-5. Epub 2017 Jan 16. Arch Virol. 2017. PMID: 28093611
-
Central Nervous System and Ocular Manifestations in Puumala Hantavirus Infection.Viruses. 2021 May 31;13(6):1040. doi: 10.3390/v13061040. Viruses. 2021. PMID: 34072819 Free PMC article. Review.
-
Puumala orthohantavirus: prevalence, biology, disease, animal models and recent advances in therapeutics development and structural biology.Front Immunol. 2025 May 8;16:1575112. doi: 10.3389/fimmu.2025.1575112. eCollection 2025. Front Immunol. 2025. PMID: 40406115 Free PMC article. Review.
Cited by
-
Genetic variants of Cao Bang hantavirus in the Chinese mole shrew (Anourosorex squamipes) and Taiwanese mole shrew (Anourosorex yamashinai).Infect Genet Evol. 2016 Jun;40:113-118. doi: 10.1016/j.meegid.2016.01.031. Epub 2016 Feb 26. Infect Genet Evol. 2016. PMID: 26921799 Free PMC article.
-
Characterization of the Puumala orthohantavirus Strains in the Northwestern Region of the Republic of Tatarstan in Relation to the Clinical Manifestations in Hemorrhagic Fever With Renal Syndrome Patients.Front Pharmacol. 2019 Sep 5;10:970. doi: 10.3389/fphar.2019.00970. eCollection 2019. Front Pharmacol. 2019. PMID: 31543819 Free PMC article.
-
Genetic Diversity of Artybash Virus in the Laxmann's Shrew (Sorex caecutiens).Vector Borne Zoonotic Dis. 2016 Jul;16(7):468-75. doi: 10.1089/vbz.2015.1903. Epub 2016 May 12. Vector Borne Zoonotic Dis. 2016. PMID: 27172519 Free PMC article.
-
Puumala Orthohantavirus Reassortant Genome Variants Likely Emerging in the Watershed Forests.Int J Mol Sci. 2023 Jan 5;24(2):1018. doi: 10.3390/ijms24021018. Int J Mol Sci. 2023. PMID: 36674534 Free PMC article.
-
Rodent-Borne Orthohantaviruses in Vietnam, Madagascar and Japan.Viruses. 2021 Jul 12;13(7):1343. doi: 10.3390/v13071343. Viruses. 2021. PMID: 34372549 Free PMC article.
References
-
- Abramson NI, Petrova TV, Dokuchaev NE, Obolenskaya EV, Lissovsky AA. Phylogeography of the gray red-backed vole Craseomys rufocanus (Rodentia: Cricetidae) across the distribution range inferred from nonrecombining molecular markers. Russian J Theriol. 2012;11:137–156.
-
- Apekina NS, Bernshtein AD, Demina VT, Gavrilovskaya IN. Variants of the immunoreactivity and infectious process in bank vole (Myodes glareolus) experimentally infected with the hantavirus Puumala (PUUV) Vopr Virusol. 2014;59:42–46. in Russian. - PubMed
-
- Asikainen K, Hanninen T, Henttonen H, Niemimaa J, Laakkonen J, Andersen HK, Bille N, Leirs H, Vaheri A, Plyusnin A. Molecular evolution of Puumala hantavirus in Fennoscandia: phylogenetic analysis of strains from two recolonization routes, Karelia and Denmark. J Gen Virol. 2000;81:2833–2841. - PubMed
-
- Dekonenko A, Yakimenko V, Ivanov A, Morozov V, Nikitin P, Khasanova S, Dzagurova T, Tkachenko E, Schmaljohn C. Genetic similarity of Puumala viruses found in Finland and western Siberia and of the mitochondrial DNA of their rodent hosts suggests a common evolutionary origin. Infect Genet Evol. 2003;3:245–257. - PubMed
-
- Dzagurova T, Tkachenko E, Slonova R, Ivanov L, Ivanidze E, Markeshin S, Dekonenko A, Niklasson B, Lundkvist A. Antigenic relationships of hantavirus strains analysed by monoclonal antibodies. Arch Virol. 1995;140:1763–1773. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

