Autophagic Degradation of the 26S Proteasome Is Mediated by the Dual ATG8/Ubiquitin Receptor RPN10 in Arabidopsis
- PMID: 26004230
- PMCID: PMC4903074
- DOI: 10.1016/j.molcel.2015.04.023
Autophagic Degradation of the 26S Proteasome Is Mediated by the Dual ATG8/Ubiquitin Receptor RPN10 in Arabidopsis
Erratum in
-
Autophagic Degradation of the 26S Proteasome Is Mediated by the Dual ATG8/Ubiquitin Receptor RPN10 in Arabidopsis.Mol Cell. 2021 May 6;81(9):2053. doi: 10.1016/j.molcel.2021.03.026. Mol Cell. 2021. PMID: 33961777 No abstract available.
Abstract
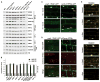
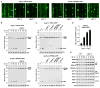
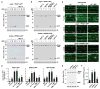
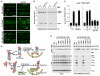
Autophagic turnover of intracellular constituents is critical for cellular housekeeping, nutrient recycling, and various aspects of growth and development in eukaryotes. Here we show that autophagy impacts the other major degradative route involving the ubiquitin-proteasome system by eliminating 26S proteasomes, a process we termed proteaphagy. Using Arabidopsis proteasomes tagged with GFP, we observed their deposition into vacuoles via a route requiring components of the autophagy machinery. This transport can be initiated separately by nitrogen starvation and chemical or genetic inhibition of the proteasome, implying distinct induction mechanisms. Proteasome inhibition stimulates comprehensive ubiquitylation of the complex, with the ensuing proteaphagy requiring the proteasome subunit RPN10, which can simultaneously bind both ATG8 and ubiquitin. Collectively, we propose that Arabidopsis RPN10 acts as a selective autophagy receptor that targets inactive 26S proteasomes by concurrent interactions with ubiquitylated proteasome subunits/targets and lipidated ATG8 lining the enveloping autophagic membranes.
Copyright © 2015 Elsevier Inc. All rights reserved.
Figures



Comment in
-
Proteaphagy-Selective Autophagy of Inactive Proteasomes.Mol Cell. 2015 Jun 18;58(6):970-1. doi: 10.1016/j.molcel.2015.06.004. Mol Cell. 2015. PMID: 26091345
References
-
- Chung T, Phillips AR, Vierstra RD. ATG8 lipidation and ATG8-mediated autophagy in Arabidopsis require ATG12 expressed from the differentially controlled ATG12A AND ATG12B loci. Plant J. 2010;62:483–493. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

