SPECTRA: An Integrated Knowledge Base for Comparing Tissue and Tumor-Specific PPI Networks in Human
- PMID: 26005672
- PMCID: PMC4424906
- DOI: 10.3389/fbioe.2015.00058
SPECTRA: An Integrated Knowledge Base for Comparing Tissue and Tumor-Specific PPI Networks in Human
Abstract
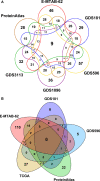
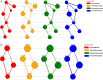
Protein-protein interaction (PPI) networks available in public repositories usually represent relationships between proteins within the cell. They ignore the specific set of tissues or tumors where the interactions take place. Indeed, proteins can form tissue-selective complexes, while they remain inactive in other tissues. For these reasons, a great attention has been recently paid to tissue-specific PPI networks, in which nodes are proteins of the global PPI network whose corresponding genes are preferentially expressed in specific tissues. In this paper, we present SPECTRA, a knowledge base to build and compare tissue or tumor-specific PPI networks. SPECTRA integrates gene expression and protein interaction data from the most authoritative online repositories. We also provide tools for visualizing and comparing such networks, in order to identify the expression and interaction changes of proteins across tissues, or between the normal and pathological states of the same tissue. SPECTRA is available as a web server at http://alpha.dmi.unict.it/spectra.
Keywords: database; interactions; proteins; tissue; tumor.
Figures







Similar articles
-
Detecting Rewiring Events in Protein-Protein Interaction Networks Based on Transcriptomic Data.Front Bioinform. 2021 Sep 8;1:724297. doi: 10.3389/fbinf.2021.724297. eCollection 2021. Front Bioinform. 2021. PMID: 36303788 Free PMC article. Review.
-
A compendium of human genes regulating feeding behavior and body weight, its functional characterization and identification of GWAS genes involved in brain-specific PPI network.BMC Genet. 2016 Dec 22;17(Suppl 3):158. doi: 10.1186/s12863-016-0466-2. BMC Genet. 2016. PMID: 28105929 Free PMC article.
-
PICKLE 2.0: A human protein-protein interaction meta-database employing data integration via genetic information ontology.PLoS One. 2017 Oct 12;12(10):e0186039. doi: 10.1371/journal.pone.0186039. eCollection 2017. PLoS One. 2017. PMID: 29023571 Free PMC article.
-
Detection of protein complexes from multiple protein interaction networks using graph embedding.Artif Intell Med. 2019 May;96:107-115. doi: 10.1016/j.artmed.2019.04.001. Epub 2019 Apr 9. Artif Intell Med. 2019. PMID: 31164203
-
Discerning molecular interactions: A comprehensive review on biomolecular interaction databases and network analysis tools.Gene. 2018 Feb 5;642:84-94. doi: 10.1016/j.gene.2017.11.028. Epub 2017 Nov 10. Gene. 2018. PMID: 29129810 Review.
Cited by
-
INBIA: a boosting methodology for proteomic network inference.BMC Bioinformatics. 2018 Jul 9;19(Suppl 7):188. doi: 10.1186/s12859-018-2183-5. BMC Bioinformatics. 2018. PMID: 30066650 Free PMC article.
-
Evolutionary diversification of the BetaM interactome acquired through co-option of the ATP1B4 gene in placental mammals.Sci Rep. 2016 Mar 4;6:22395. doi: 10.1038/srep22395. Sci Rep. 2016. PMID: 26939788 Free PMC article.
-
The TissueNet v.2 database: A quantitative view of protein-protein interactions across human tissues.Nucleic Acids Res. 2017 Jan 4;45(D1):D427-D431. doi: 10.1093/nar/gkw1088. Epub 2016 Nov 29. Nucleic Acids Res. 2017. PMID: 27899616 Free PMC article.
-
Detecting Rewiring Events in Protein-Protein Interaction Networks Based on Transcriptomic Data.Front Bioinform. 2021 Sep 8;1:724297. doi: 10.3389/fbinf.2021.724297. eCollection 2021. Front Bioinform. 2021. PMID: 36303788 Free PMC article. Review.
-
PPIXpress and PPICompare webservers infer condition-specific and differential PPI networks.Bioinform Adv. 2025 Feb 11;5(1):vbaf003. doi: 10.1093/bioadv/vbaf003. eCollection 2025. Bioinform Adv. 2025. PMID: 39990257 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

