Development of genetic tools for Myceliophthora thermophila
- PMID: 26013561
- PMCID: PMC4446196
- DOI: 10.1186/s12896-015-0165-5
Development of genetic tools for Myceliophthora thermophila
Abstract
Background: The thermophilic filamentous fungus Myceliophthora thermophila has many suitable characteristics for industrial biotechnology and could be a promising new chassis system for synthetic biology, particularly the ATCC 42464 strain, whose genome was sequenced in 2011. However, metabolic engineering of this strain using genetic approaches has not been reported owing to a lack of genetic tools for this organism.
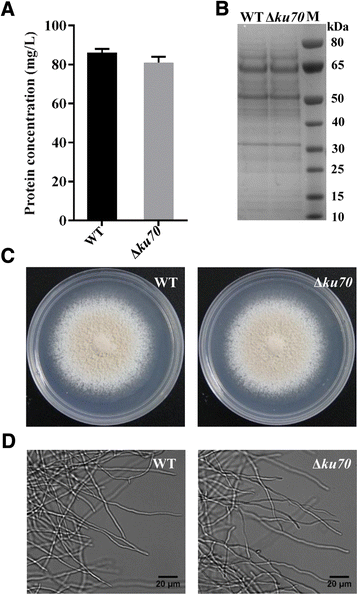
Results: In the present study, we developed a high efficiency Agrobacterium tumefaciens mediated transformation system for M. thermophila, including an approach for targeted gene deletion using green fluorescence protein (GFP) as a marker for selection. Up to 145 transformants per 10(5) conidia were obtained in one transformation plate. Moreover, a ku70 deletion mutant was constructed in the ATCC 42464 background using the tools developed in present study and subsequently characterized. The ku70 deletion construct was designed using resistance to phosphinothricin as the selection marker. Additionally, a GFP-encoding cassette was incorporated that allowed for the selection of site-specific (no fluorescence) or ectopic (fluorescence) integration of the ku70 construct. Transformants with ectopically integrated ku70 deletion constructs were therefore identified using the fluorescent signal of GFP. PCR and Southern blotting analyses of non-fluorescent putative ku70 deletion transformants revealed all 11 tested transformants to be correct deletions. The deletion frequency in a pool of 116 transformants analyzed was 58 %. Moreover, the homologous rate improved about 3 folds under ku70 mutant using the pyrG as a test gene to disrupt in M. thermophila.
Conclusions: We successfully developed an efficient transformation and target gene disruption approach for M. thermophila ATCC 42464 mediated by A. tumefaciens. The tools and the ku70 deletion strain developed here should advance the development of M. thermophila as an industrial host through metabolic engineering and accelerate the elucidation of the mechanism of rapid cellulose degradation in this thermophilic fungus.
Figures





Similar articles
-
Highly efficient generation of signal transduction knockout mutants using a fungal strain deficient in the mammalian ku70 ortholog.Gene. 2006 Aug 15;378:1-10. doi: 10.1016/j.gene.2006.03.020. Epub 2006 Apr 30. Gene. 2006. PMID: 16814491
-
Molecular characterization of KU70 and KU80 homologues and exploitation of a KU70-deficient mutant for improving gene deletion frequency in Rhodosporidium toruloides.BMC Microbiol. 2014 Feb 27;14:50. doi: 10.1186/1471-2180-14-50. BMC Microbiol. 2014. PMID: 25188820 Free PMC article.
-
A ku70 null mutant improves gene targeting frequency in the fungal pathogen Verticillium dahliae.World J Microbiol Biotechnol. 2015 Dec;31(12):1889-97. doi: 10.1007/s11274-015-1907-1. World J Microbiol Biotechnol. 2015. PMID: 26475327
-
Myceliophthora thermophila as promising fungal cell factories for industrial bioproduction: From rational design to industrial applications.Bioresour Technol. 2025 Mar;419:132051. doi: 10.1016/j.biortech.2025.132051. Epub 2025 Jan 9. Bioresour Technol. 2025. PMID: 39798815 Review.
-
Approaches to genetic tool development for rapid domestication of non-model microorganisms.Biotechnol Biofuels. 2021 Jan 25;14(1):30. doi: 10.1186/s13068-020-01872-z. Biotechnol Biofuels. 2021. PMID: 33494801 Free PMC article. Review.
Cited by
-
Dissecting cellobiose metabolic pathway and its application in biorefinery through consolidated bioprocessing in Myceliophthora thermophila.Fungal Biol Biotechnol. 2019 Nov 13;6:21. doi: 10.1186/s40694-019-0083-8. eCollection 2019. Fungal Biol Biotechnol. 2019. PMID: 31754437 Free PMC article.
-
Engineered Fungus Thermothelomyces thermophilus Producing Plant Storage Proteins.J Fungi (Basel). 2022 Jan 26;8(2):119. doi: 10.3390/jof8020119. J Fungi (Basel). 2022. PMID: 35205873 Free PMC article.
-
Metabolic engineering of the cellulolytic thermophilic fungus Myceliophthora thermophila to produce ethanol from cellobiose.Biotechnol Biofuels. 2020 Feb 1;13:23. doi: 10.1186/s13068-020-1661-y. eCollection 2020. Biotechnol Biofuels. 2020. PMID: 32021654 Free PMC article.
-
Coordination of consolidated bioprocessing technology and carbon dioxide fixation to produce malic acid directly from plant biomass in Myceliophthora thermophila.Biotechnol Biofuels. 2021 Sep 23;14(1):186. doi: 10.1186/s13068-021-02042-5. Biotechnol Biofuels. 2021. PMID: 34556173 Free PMC article.
-
Characterizing the Role of AosfgA and AofluG in Mycelial and Conidial Development in Arthrobotrys oligospora and Their Role in Secondary Metabolism.Microorganisms. 2024 Mar 19;12(3):615. doi: 10.3390/microorganisms12030615. Microorganisms. 2024. PMID: 38543666 Free PMC article.
References
-
- Singh B. Myceliophthora thermophila syn. Sporotrichum thermophile: a thermophilic mould of biotechnological potential. Crit Rev Biotechnol 2014; 15: 1-11. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

