Different STAT Transcription Complexes Drive Early and Delayed Responses to Type I IFNs
- PMID: 26019270
- PMCID: PMC4617678
- DOI: 10.4049/jimmunol.1401139
Different STAT Transcription Complexes Drive Early and Delayed Responses to Type I IFNs
Abstract
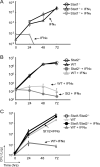
IFNs, which transduce pivotal signals through Stat1 and Stat2, effectively suppress the replication of Legionella pneumophila in primary murine macrophages. Although the ability of IFN-γ to impede L. pneumophila growth is fully dependent on Stat1, IFN-αβ unexpectedly suppresses L. pneumophila growth in both Stat1- and Stat2-deficient macrophages. New studies demonstrating that the robust response to IFN-αβ is lost in Stat1-Stat2 double-knockout macrophages suggest that Stat1 and Stat2 are functionally redundant in their ability to direct an innate response toward L. pneumophila. Because the ability of IFN-αβ to signal through Stat1-dependent complexes (i.e., Stat1-Stat1 and Stat1-Stat2 dimers) has been well characterized, the current studies focus on how Stat2 is able to direct a potent response to IFN-αβ in the absence of Stat1. These studies reveal that IFN-αβ is able to drive the formation of a Stat2 and IFN regulatory factor 9 complex that drives the expression of a subset of IFN-stimulated genes, but with substantially delayed kinetics. These observations raise the possibility that this pathway evolved in response to microbes that have devised strategies to subvert Stat1-dependent responses.
Copyright © 2015 by The American Association of Immunologists, Inc.
Figures




References
-
- Mykietiuk A, Carratala J, Fernandez-Sabe N, Dorca J, Verdaguer R, Manresa F, Gudiol F. Clinical outcomes for hospitalized patients with Legionella pneumonia in the antigenuria era: the influence of levofloxacin therapy. Clin Infect Dis. 2005;40:794–799. - PubMed
-
- Segal G, Feldman M, Zusman T. The Icm/Dot type-IV secretion systems of Legionella pneumophila and Coxiella burnetii. FEMS Microbiol Rev. 2005;29:65–81. - PubMed
-
- Schindler C, Levy DE, Decker T. JAK-STAT signaling: from interferons to cytokines. The Journal of biological chemistry. 2007;282:20059–20063. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

