Revisiting molecular serotyping of Streptococcus pneumoniae
- PMID: 26041622
- PMCID: PMC4460616
- DOI: 10.1186/1471-2164-16-S5-S1
Revisiting molecular serotyping of Streptococcus pneumoniae
Abstract
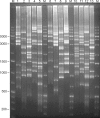
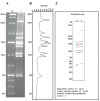
Background: Ninety-two Streptococcus pneumoniae serotypes have been described so far, but the pneumococcal conjugate vaccine introduced in the Brazilian basic vaccination schedule in 2010 covers only the ten most prevalent in the country. Pneumococcal serotype-shifting after massive immunization is a major concern and monitoring this phenomenon requires efficient and accessible serotyping methods. Pneumococcal serotyping based on antisera produced in animals is laborious and restricted to a few reference laboratories. Alternatively, molecular serotyping methods assess polymorphisms in the cps gene cluster, which encodes key enzymes for capsular polysaccharides synthesis in pneumococci. In one such approach, cps-RFLP, the PCR amplified cps loci are digested with an endonuclease, generating serotype-specific fingerprints on agarose gel electrophoresis.
Methods: In this work, in silico and in vitro approaches were combined to demonstrate that XhoII is the most discriminating endonuclease for cps-RFLP, and to build a database of serotype-specific fingerprints that accommodates the genetic diversity within the cps locus of 92 known pneumococci serotypes.
Results: The expected specificity of cps-RFLP using XhoII was 76% for serotyping and 100% for serogrouping. The database of cps-RFLP fingerprints was integrated to Molecular Serotyping Tool (MST), a previously published web-based software for molecular serotyping. In addition, 43 isolates representing 29 serotypes prevalent in the state of Minas Gerais, Brazil, from 2007 to 2013, were examined in vitro; 11 serotypes (nine serogroups) matched the respective in silico patterns calculated for reference strains. The remaining experimental patterns, despite their resemblance to their expected in silico patterns, did not reach the threshold of similarity score to be considered a match and were then added to the database.
Conclusion: The cps-RFLP method with XhoII outperformed the antisera-based and other molecular serotyping methods in regard of the expected specificity. In order to accommodate the genetic variability of the pneumococci cps loci, the database of cps-RFLP patterns will be progressively expanded to include new variant in vitro patterns. The cps-RFLP method with endonuclease XhoII coupled with MST for computer-assisted interpretation of results may represent a relevant contribution to the real time detection of changes in regional pneumococci population diversity in response to mass immunization programs.
Figures


References
-
- O'Brien KL, Wolfson LJ, Watt JP, Henkle E, Deloria-Knoll M, McCall N, Lee E, Mulholland K, Levine OS, Cherian T. et al.Burden of disease caused by Streptococcus pneumoniae in children younger than 5 years: global estimates. Lancet. 2009;374(9693):893–902. doi: 10.1016/S0140-6736(09)61204-6. - DOI - PubMed
-
- Pichon B, Ladhani SN, Slack MP, Segonds-Pichon A, Andrews NJ, Waight PA, Miller E, George R. Changes in molecular epidemiology of streptococcus pneumoniae causing meningitis following introduction of pneumococcal conjugate vaccination in England and Wales. J Clin Microbiol. 2013;51(3):820–827. doi: 10.1128/JCM.01917-12. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

