DT-Web: a web-based application for drug-target interaction and drug combination prediction through domain-tuned network-based inference
- PMID: 26050742
- PMCID: PMC4464606
- DOI: 10.1186/1752-0509-9-S3-S4
DT-Web: a web-based application for drug-target interaction and drug combination prediction through domain-tuned network-based inference
Abstract
Background: The identification of drug-target interactions (DTI) is a costly and time-consuming step in drug discovery and design. Computational methods capable of predicting reliable DTI play an important role in the field. Algorithms may aim to design new therapies based on a single approved drug or a combination of them. Recently, recommendation methods relying on network-based inference in connection with knowledge coming from the specific domain have been proposed.
Description: Here we propose a web-based interface to the DT-Hybrid algorithm, which applies a recommendation technique based on bipartite network projection implementing resources transfer within the network. This technique combined with domain-specific knowledge expressing drugs and targets similarity is used to compute recommendations for each drug. Our web interface allows the users: (i) to browse all the predictions inferred by the algorithm; (ii) to upload their custom data on which they wish to obtain a prediction through a DT-Hybrid based pipeline; (iii) to help in the early stages of drug combinations, repositioning, substitution, or resistance studies by finding drugs that can act simultaneously on multiple targets in a multi-pathway environment. Our system is periodically synchronized with DrugBank and updated accordingly. The website is free, open to all users, and available at http://alpha.dmi.unict.it/dtweb/.
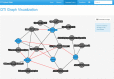
Conclusions: Our web interface allows users to search and visualize information on drugs and targets eventually providing their own data to compute a list of predictions. The user can visualize information about the characteristics of each drug, a list of predicted and validated targets, associated enzymes and transporters. A table containing key information and GO classification allows the users to perform their own analysis on our data. A special interface for data submission allows the execution of a pipeline, based on DT-Hybrid, predicting new targets with the corresponding p-values expressing the reliability of each group of predictions. Finally, It is also possible to specify a list of genes tracking down all the drugs that may have an indirect influence on them based on a multi-drug, multi-target, multi-pathway analysis, which aims to discover drugs for future follow-up studies.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

