TRIM32 Senses and Restricts Influenza A Virus by Ubiquitination of PB1 Polymerase
- PMID: 26057645
- PMCID: PMC4461266
- DOI: 10.1371/journal.ppat.1004960
TRIM32 Senses and Restricts Influenza A Virus by Ubiquitination of PB1 Polymerase
Abstract
Polymerase basic protein 1 (PB1) is the catalytic core of the influenza A virus (IAV) RNA polymerase complex essential for viral transcription and replication. Understanding the intrinsic mechanisms which block PB1 function could stimulate development of new anti-influenza therapeutics. Affinity purification coupled with mass spectrometry (AP-MS) was used to identify host factors interacting with PB1. Among PB1 interactors, the E3 ubiquitin ligase TRIM32 interacts with PB1 proteins derived from multiple IAV strains. TRIM32 senses IAV infection by interacting with PB1 and translocates with PB1 to the nucleus following influenza infection. Ectopic TRIM32 expression attenuates IAV infection. Conversely, RNAi depletion and knockout of TRIM32 increase susceptibility of tracheal and lung epithelial cells to IAV infection. Reconstitution of trim32-/- mouse embryonic fibroblasts with TRIM32, but not a catalytically inactive mutant, restores viral restriction. Furthermore, TRIM32 directly ubiquitinates PB1, leading to PB1 protein degradation and subsequent reduction of polymerase activity. Thus, TRIM32 is an intrinsic IAV restriction factor which senses and targets the PB1 polymerase for ubiquitination and protein degradation. TRIM32 represents a model of intrinsic immunity, in which a host protein directly senses and counters viral infection in a species specific fashion by directly limiting viral replication.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



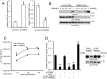



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

