A survey on the computational approaches to identify drug targets in the postgenomic era
- PMID: 26060814
- PMCID: PMC4427773
- DOI: 10.1155/2015/239654
A survey on the computational approaches to identify drug targets in the postgenomic era
Abstract
Identifying drug targets plays essential roles in designing new drugs and combating diseases. Unfortunately, our current knowledge about drug targets is far from comprehensive. Screening drug targets in the lab is an expensive and time-consuming procedure. In the past decade, the accumulation of various types of omics data makes it possible to develop computational approaches to predict drug targets. In this paper, we make a survey on the recent progress being made on computational methodologies that have been developed to predict drug targets based on different kinds of omics data and drug property data.
Figures
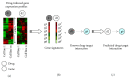


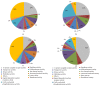
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

