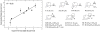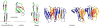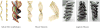Membrane Protein Structure, Function, and Dynamics: a Perspective from Experiments and Theory
- PMID: 26063070
- PMCID: PMC4515176
- DOI: 10.1007/s00232-015-9802-0
Membrane Protein Structure, Function, and Dynamics: a Perspective from Experiments and Theory
Abstract
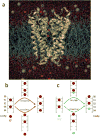
Membrane proteins mediate processes that are fundamental for the flourishing of biological cells. Membrane-embedded transporters move ions and larger solutes across membranes; receptors mediate communication between the cell and its environment and membrane-embedded enzymes catalyze chemical reactions. Understanding these mechanisms of action requires knowledge of how the proteins couple to their fluid, hydrated lipid membrane environment. We present here current studies in computational and experimental membrane protein biophysics, and show how they address outstanding challenges in understanding the complex environmental effects on the structure, function, and dynamics of membrane proteins.
Figures