Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation
- PMID: 26089354
- PMCID: PMC4845908
- DOI: 10.1126/science.aab2276
Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation
Abstract
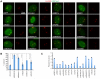
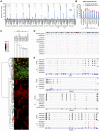
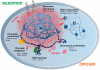
The inactive X chromosome (Xi) serves as a model to understand gene silencing on a global scale. Here, we perform "identification of direct RNA interacting proteins" (iDRiP) to isolate a comprehensive protein interactome for Xist, an RNA required for Xi silencing. We discover multiple classes of interactors-including cohesins, condensins, topoisomerases, RNA helicases, chromatin remodelers, and modifiers-that synergistically repress Xi transcription. Inhibiting two or three interactors destabilizes silencing. Although Xist attracts some interactors, it repels architectural factors. Xist evicts cohesins from the Xi and directs an Xi-specific chromosome conformation. Upon deleting Xist, the Xi acquires the cohesin-binding and chromosomal architecture of the active X. Our study unveils many layers of Xi repression and demonstrates a central role for RNA in the topological organization of mammalian chromosomes.
Copyright © 2015, American Association for the Advancement of Science.
Figures



References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

