FOXM1 binds directly to non-consensus sequences in the human genome
- PMID: 26100407
- PMCID: PMC4492089
- DOI: 10.1186/s13059-015-0696-z
FOXM1 binds directly to non-consensus sequences in the human genome
Abstract
Background: The Forkhead (FKH) transcription factor FOXM1 is a key regulator of the cell cycle and is overexpressed in most types of cancer. FOXM1, similar to other FKH factors, binds to a canonical FKH motif in vitro. However, genome-wide mapping studies in different cell lines have shown a lack of enrichment of the FKH motif, suggesting an alternative mode of chromatin recruitment. We have investigated the role of direct versus indirect DNA binding in FOXM1 recruitment by performing ChIP-seq with wild-type and DNA binding deficient FOXM1.
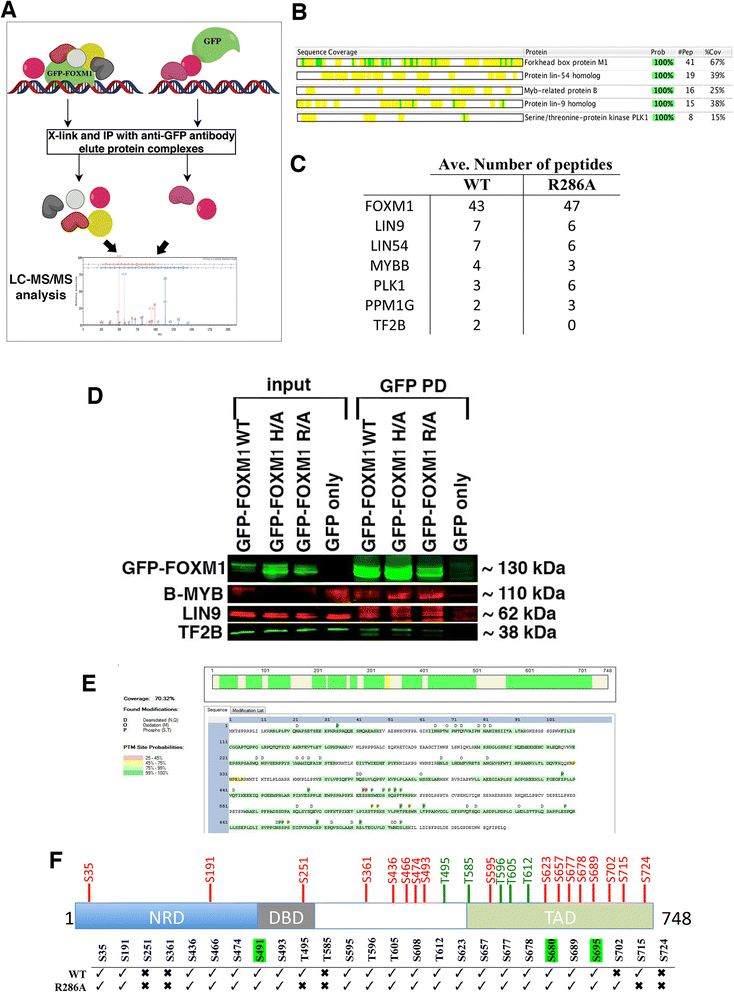
Results: An in vitro fluorescence polarization assay identified point mutations in the DNA binding domain of FOXM1 that inhibit binding to a FKH consensus sequence. Cell lines expressing either wild-type or DNA binding deficient GFP-tagged FOXM1 were used for genome-wide mapping studies comparing the distribution of the DNA binding deficient protein to the wild-type. This shows that interaction of the FOXM1 DNA binding domain with target DNA is essential for recruitment. Moreover, analysis of the protein interactome of wild-type versus DNA binding deficient FOXM1 shows that the reduced recruitment is not due to inhibition of protein-protein interactions.
Conclusions: A functional DNA binding domain is essential for FOXM1 chromatin recruitment. Even in FOXM1 mutants with almost complete loss of binding, the protein-protein interactions and pattern of phosphorylation are largely unaffected. These results strongly support a model whereby FOXM1 is specifically recruited to chromatin through co-factor interactions by binding directly to non-canonical DNA sequences.
Figures






References
-
- Teh MT, Wong ST, Neill GW, Ghali LR, Philpott MP, Quinn AG. FOXM1 is a downstream target of Gli1 in basal cell carcinomas. Cancer Res. 2002;62:4773–4780. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

