ECHO-liveFISH: in vivo RNA labeling reveals dynamic regulation of nuclear RNA foci in living tissues
- PMID: 26101260
- PMCID: PMC4627062
- DOI: 10.1093/nar/gkv614
ECHO-liveFISH: in vivo RNA labeling reveals dynamic regulation of nuclear RNA foci in living tissues
Abstract
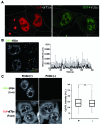
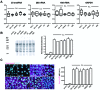
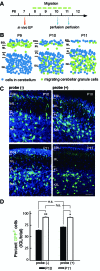
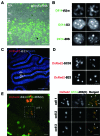
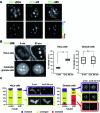
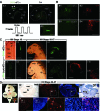
Elucidating the dynamic organization of nuclear RNA foci is important for understanding and manipulating these functional sites of gene expression in both physiological and pathological states. However, such studies have been difficult to establish in vivo as a result of the absence of suitable RNA imaging methods. Here, we describe a high-resolution fluorescence RNA imaging method, ECHO-liveFISH, to label endogenous nuclear RNA in living mice and chicks. Upon in vivo electroporation, exciton-controlled sequence-specific oligonucleotide probes revealed focally concentrated endogenous 28S rRNA and U3 snoRNA at nucleoli and poly(A) RNA at nuclear speckles. Time-lapse imaging reveals steady-state stability of these RNA foci and dynamic dissipation of 28S rRNA concentrations upon polymerase I inhibition in native brain tissue. Confirming the validity of this technique in a physiological context, the in vivo RNA labeling did not interfere with the function of target RNA nor cause noticeable cytotoxicity or perturbation of cellular behavior.
© The Author(s) 2015. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


Similar articles
-
Live Imaging of Nuclear RNPs in Mammalian Complex Tissue with ECHO-liveFISH.Methods Mol Biol. 2018;1649:259-272. doi: 10.1007/978-1-4939-7213-5_17. Methods Mol Biol. 2018. PMID: 29130203
-
In situ hybridization in living cells: detection of RNA molecules.Exp Cell Res. 1997 Feb 25;231(1):226-33. doi: 10.1006/excr.1996.3464. Exp Cell Res. 1997. PMID: 9056430
-
Nanoscale imaging of RNA with expansion microscopy.Nat Methods. 2016 Aug;13(8):679-84. doi: 10.1038/nmeth.3899. Epub 2016 Jul 4. Nat Methods. 2016. PMID: 27376770 Free PMC article.
-
Visualizing RNA molecules inside the nucleus of living cells.Methods. 2003 Jan;29(1):51-7. doi: 10.1016/s1046-2023(02)00290-6. Methods. 2003. PMID: 12543071 Review.
-
Nanostructured probes for RNA detection in living cells.Ann Biomed Eng. 2006 Jan;34(1):39-50. doi: 10.1007/s10439-005-9003-6. Epub 2006 Feb 7. Ann Biomed Eng. 2006. PMID: 16463087 Review.
Cited by
-
Subcellular Transcriptomics and Proteomics: A Comparative Methods Review.Mol Cell Proteomics. 2022 Feb;21(2):100186. doi: 10.1016/j.mcpro.2021.100186. Epub 2021 Dec 16. Mol Cell Proteomics. 2022. PMID: 34922010 Free PMC article. Review.
-
DNA/RNA Fluorescence Imaging by Synthetic Nucleic Acids.Adv Exp Med Biol. 2021;1310:475-493. doi: 10.1007/978-981-33-6064-8_17. Adv Exp Med Biol. 2021. PMID: 33834446
-
Oligonucleotide Probe for Transcriptome in Vivo Analysis (TIVA) of Single Neurons with Minimal Background.ACS Chem Biol. 2020 Oct 16;15(10):2714-2721. doi: 10.1021/acschembio.0c00499. Epub 2020 Sep 23. ACS Chem Biol. 2020. PMID: 32902259 Free PMC article.
-
Design of thiazole orange oligonucleotide probes for detection of DNA and RNA by fluorescence and duplex melting.Org Biomol Chem. 2019 Jun 28;17(24):5943-5950. doi: 10.1039/c9ob00885c. Epub 2019 Jun 3. Org Biomol Chem. 2019. PMID: 31157811 Free PMC article.
-
Real-time imaging of RNA polymerase I activity in living human cells.J Cell Biol. 2023 Jan 2;222(1):e202202110. doi: 10.1083/jcb.202202110. Epub 2022 Oct 25. J Cell Biol. 2023. PMID: 36282216 Free PMC article.
References
-
- Lewis J.D., Tollervey D. Like attracts like: getting RNA processing together in the nucleus. Science. 2000;288:1385–1389. - PubMed
-
- Cioce M., Lamond A.I. Cajal bodies: a long history of discovery. Annu. Rev. Cell Dev. Biol. 2005;21:105–131. - PubMed
-
- Gubitz A.K., Feng W., Dreyfuss G. The SMN complex. Exp. Cell Res. 2004;296:51–56. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

