Phylodynamics of H5N1 Highly Pathogenic Avian Influenza in Europe, 2005-2010: Potential for Molecular Surveillance of New Outbreaks
- PMID: 26110587
- PMCID: PMC4488740
- DOI: 10.3390/v7062773
Phylodynamics of H5N1 Highly Pathogenic Avian Influenza in Europe, 2005-2010: Potential for Molecular Surveillance of New Outbreaks
Abstract
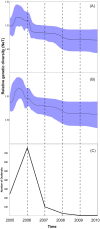
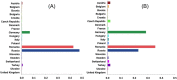
Previous Bayesian phylogeographic studies of H5N1 highly pathogenic avian influenza viruses (HPAIVs) explored the origin and spread of the epidemic from China into Russia, indicating that HPAIV circulated in Russia prior to its detection there in 2005. In this study, we extend this research to explore the evolution and spread of HPAIV within Europe during the 2005-2010 epidemic, using all available sequences of the hemagglutinin (HA) and neuraminidase (NA) gene regions that were collected in Europe and Russia during the outbreak. We use discrete-trait phylodynamic models within a Bayesian statistical framework to explore the evolution of HPAIV. Our results indicate that the genetic diversity and effective population size of HPAIV peaked between mid-2005 and early 2006, followed by drastic decline in 2007, which coincides with the end of the epidemic in Europe. Our results also suggest that domestic birds were the most likely source of the spread of the virus from Russia into Europe. Additionally, estimates of viral dispersal routes indicate that Russia, Romania, and Germany were key epicenters of these outbreaks. Our study quantifies the dynamics of a major European HPAIV pandemic and substantiates the ability of phylodynamic models to improve molecular surveillance of novel AIVs.
Keywords: Bayesian inference; Europe; H5N1; Russia; highly pathogenic avian influenza; phylodynamic models; phylogeography; surveillance.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

