Decreased Expression of CoREST1 and CoREST2 Together with LSD1 and HDAC1/2 during Neuronal Differentiation
- PMID: 26111147
- PMCID: PMC4482511
- DOI: 10.1371/journal.pone.0131760
Decreased Expression of CoREST1 and CoREST2 Together with LSD1 and HDAC1/2 during Neuronal Differentiation
Erratum in
-
Correction: Decreased Expression of CoREST1 and CoREST2 Together with LSD1 and HDAC1/2 during Neuronal Differentiation.PLoS One. 2015 Jul 17;10(7):e0133555. doi: 10.1371/journal.pone.0133555. eCollection 2015. PLoS One. 2015. PMID: 26186538 Free PMC article. No abstract available.
Abstract
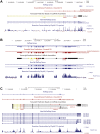
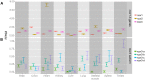
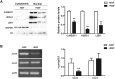
CoREST (CoREST1, rcor1) transcriptional corepressor together with the histone demethylase LSD1 (KDM1A) and the histone deacetylases HDAC1/2 form LSD1-CoREST-HDAC (LCH) transcriptional complexes to regulate gene expression. CoREST1 belong to a family that also comprises CoREST2 (rcor2) and CoREST3 (rcor3). CoREST1 represses the expression of neuronal genes during neuronal differentiation. However, the role of paralogs CoREST2 and CoREST3 in this process is just starting to emerge. Here, we report the expression of all CoRESTs and partners LSD1 and HDAC1/2 in two models of neuronal differentiation: Nerve-Growth-Factor (NGF)-induced neuronal phenotype of PC12 cells, and in vitro maturation of embryonic rat cortical neurons. In both models, a concomitant and gradual decrease of LSD1, HDAC1, HDAC2, CoREST1, and CoREST2, but not CoREST3 was observed. As required by the study, full-length rat rcor1 gene was identified using in silico analysis of available rat genome. The work was also complemented by the analysis of rat RNA-seq databases. The analysis showed that all CoRESTs, including the identified four splicing variants of rat CoREST3, display a wide expression in adult tissues. Moreover, the analysis of RNA-seq databases showed that CoREST2 displays a higher expression than CoREST1 and CoREST3 in the mature brain. Immunofluorescent assays and immunoblots of adult rat brain showed that all CoRESTs are present in both glia and neurons. Regarding functional partnership, CoREST2 and CoREST3 interact with all LSD1 splicing variants. In conclusion, neuronal differentiation is accompanied by decreased expression of all core components of LCH complexes, but not CoREST3. The combination of the differential transcriptional repressor capacity of LCH complexes and variable protein levels of its different components should result in a finely tuned gene expression during neuronal differentiation and in the adult brain.
Conflict of interest statement
Figures

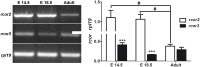


Similar articles
-
Differential properties of transcriptional complexes formed by the CoREST family.Mol Cell Biol. 2014 Jul;34(14):2760-70. doi: 10.1128/MCB.00083-14. Mol Cell Biol. 2014. PMID: 24820421 Free PMC article.
-
PIASγ controls stability and facilitates SUMO-2 conjugation to CoREST family of transcriptional co-repressors.Biochem J. 2018 Apr 23;475(8):1441-1454. doi: 10.1042/BCJ20170983. Biochem J. 2018. PMID: 29555846
-
Phosphorylation of neuronal Lysine-Specific Demethylase 1LSD1/KDM1A impairs transcriptional repression by regulating interaction with CoREST and histone deacetylases HDAC1/2.J Neurochem. 2014 Mar;128(5):603-16. doi: 10.1111/jnc.12457. Epub 2013 Oct 23. J Neurochem. 2014. PMID: 24111946
-
More than a Corepressor: The Role of CoREST Proteins in Neurodevelopment.eNeuro. 2020 Mar 10;7(2):ENEURO.0337-19.2020. doi: 10.1523/ENEURO.0337-19.2020. Print 2020 Mar/Apr. eNeuro. 2020. PMID: 32075869 Free PMC article. Review.
-
Histone deacetylases 1 and 2 are required for brain development.Int J Dev Biol. 2015;59(4-6):171-7. doi: 10.1387/ijdb.150071tz. Int J Dev Biol. 2015. PMID: 26198144 Review.
Cited by
-
Correction: Decreased Expression of CoREST1 and CoREST2 Together with LSD1 and HDAC1/2 during Neuronal Differentiation.PLoS One. 2015 Jul 17;10(7):e0133555. doi: 10.1371/journal.pone.0133555. eCollection 2015. PLoS One. 2015. PMID: 26186538 Free PMC article. No abstract available.
-
Revealing RCOR2 as a regulatory component of nuclear speckles.Epigenetics Chromatin. 2021 Nov 24;14(1):51. doi: 10.1186/s13072-021-00425-4. Epigenetics Chromatin. 2021. PMID: 34819154 Free PMC article.
-
Role of primary aging hallmarks in Alzheimer´s disease.Theranostics. 2023 Jan 1;13(1):197-230. doi: 10.7150/thno.79535. eCollection 2023. Theranostics. 2023. PMID: 36593969 Free PMC article. Review.
-
Biological and therapeutic role of LSD1 in Alzheimer's diseases.Front Pharmacol. 2022 Oct 25;13:1020556. doi: 10.3389/fphar.2022.1020556. eCollection 2022. Front Pharmacol. 2022. PMID: 36386192 Free PMC article. Review.
-
Coordinated regulation of the ribosome and proteasome by PRMT1 in the maintenance of neural stemness in cancer cells and neural stem cells.J Biol Chem. 2021 Nov;297(5):101275. doi: 10.1016/j.jbc.2021.101275. Epub 2021 Oct 4. J Biol Chem. 2021. PMID: 34619150 Free PMC article.
References
-
- Humphrey GW, Wang Y, Russanova VR, Hirai T, Qin J, Nakatani Y, et al. (2001) Stable histone deacetylase complexes distinguished by the presence of SANT domain proteins CoREST/kiaa0071 and Mta-L1. J Biol Chem 276: 6817–6824. - PubMed
-
- Zibetti C, Adamo A, Binda C, Forneris F, Toffolo E, Verpelli C, et al. (2010) Alternative splicing of the histone demethylase LSD1/KDM1 contributes to the modulation of neurite morphogenesis in the mammalian nervous system. J Neurosci 30: 2521–2532. 10.1523/JNEUROSCI.5500-09.2010 - DOI - PMC - PubMed
-
- Yang M, Gocke CB, Luo X, Borek D, Tomchick DR, Machius M, et al. (2006) Structural basis for CoREST-dependent demethylation of nucleosomes by the human LSD1 histone demethylase. Mol Cell 23: 377–387. - PubMed
-
- Forneris F, Binda C, Adamo A, Battaglioli E, Mattevi A (2007) Structural basis of LSD1-CoREST selectivity in histone H3 recognition. J Biol Chem 282: 20070–20074. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

