Surfing the Protein-Protein Interaction Surface Using Docking Methods: Application to the Design of PPI Inhibitors
- PMID: 26111183
- PMCID: PMC6272567
- DOI: 10.3390/molecules200611569
Surfing the Protein-Protein Interaction Surface Using Docking Methods: Application to the Design of PPI Inhibitors
Abstract
Blocking protein-protein interactions (PPI) using small molecules or peptides modulates biochemical pathways and has therapeutic significance. PPI inhibition for designing drug-like molecules is a new area that has been explored extensively during the last decade. Considering the number of available PPI inhibitor databases and the limited number of 3D structures available for proteins, docking and scoring methods play a major role in designing PPI inhibitors as well as stabilizers. Docking methods are used in the design of PPI inhibitors at several stages of finding a lead compound, including modeling the protein complex, screening for hot spots on the protein-protein interaction interface and screening small molecules or peptides that bind to the PPI interface. There are three major challenges to the use of docking on the relatively flat surfaces of PPI. In this review we will provide some examples of the use of docking in PPI inhibitor design as well as its limitations. The combination of experimental and docking methods with improved scoring function has thus far resulted in few success stories of PPI inhibitors for therapeutic purposes. Docking algorithms used for PPI are in the early stages, however, and as more data are available docking will become a highly promising area in the design of PPI inhibitors or stabilizers.
Keywords: docking; drug-like molecules; hot spots; protein docking; protein-protein interactions; virtual screening.
Conflict of interest statement
The authors declare no conflict of interest.
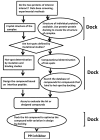
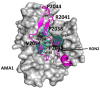
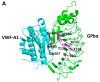
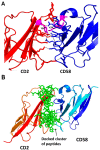
Figures

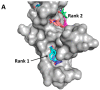

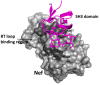
Similar articles
-
In silico structure-based approaches to discover protein-protein interaction-targeting drugs.Methods. 2017 Dec 1;131:22-32. doi: 10.1016/j.ymeth.2017.08.006. Epub 2017 Aug 9. Methods. 2017. PMID: 28802714 Free PMC article. Review.
-
Fast Rescoring Protocols to Improve the Performance of Structure-Based Virtual Screening Performed on Protein-Protein Interfaces.J Chem Inf Model. 2020 Aug 24;60(8):3910-3934. doi: 10.1021/acs.jcim.0c00545. Epub 2020 Aug 11. J Chem Inf Model. 2020. PMID: 32786511
-
A novel small-molecule PPI inhibitor targeting integrin αvβ3-osteopontin interface blocks bone resorption in vitro and prevents bone loss in mice.Biomaterials. 2016 Aug;98:131-42. doi: 10.1016/j.biomaterials.2016.05.007. Epub 2016 May 5. Biomaterials. 2016. PMID: 27187277
-
Benchmark Study Based on 2P2IDB to Gain Insights into the Discovery of Small-Molecule PPI Inhibitors.J Phys Chem B. 2018 Mar 8;122(9):2544-2555. doi: 10.1021/acs.jpcb.7b12658. Epub 2018 Feb 22. J Phys Chem B. 2018. PMID: 29420886
-
State-of-the-art strategies for targeting protein-protein interactions by small-molecule inhibitors.Chem Soc Rev. 2015 Nov 21;44(22):8238-59. doi: 10.1039/c5cs00252d. Epub 2015 Aug 6. Chem Soc Rev. 2015. PMID: 26248294 Review.
Cited by
-
Molecular docking as a popular tool in drug design, an in silico travel.Adv Appl Bioinform Chem. 2016 Jun 28;9:1-11. doi: 10.2147/AABC.S105289. eCollection 2016. Adv Appl Bioinform Chem. 2016. PMID: 27390530 Free PMC article. Review.
-
Design and Synthesis of Novel Helix Mimetics Based on the Covalent H-Bond Replacement and Amide Surrogate.Molecules. 2023 Jan 12;28(2):780. doi: 10.3390/molecules28020780. Molecules. 2023. PMID: 36677838 Free PMC article.
-
Current Challenges and Opportunities in Designing Protein-Protein Interaction Targeted Drugs.Adv Appl Bioinform Chem. 2020 Nov 12;13:11-25. doi: 10.2147/AABC.S235542. eCollection 2020. Adv Appl Bioinform Chem. 2020. PMID: 33209039 Free PMC article. Review.
-
Molecular Dynamics and Other HPC Simulations for Drug Discovery.Methods Mol Biol. 2024;2716:265-291. doi: 10.1007/978-1-0716-3449-3_12. Methods Mol Biol. 2024. PMID: 37702944
-
Interfacial Peptides as Affinity Modulating Agents of Protein-Protein Interactions.Biomolecules. 2022 Jan 8;12(1):106. doi: 10.3390/biom12010106. Biomolecules. 2022. PMID: 35053254 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

