Whole-genome sequencing of uropathogenic Escherichia coli reveals long evolutionary history of diversity and virulence
- PMID: 26112070
- PMCID: PMC4530057
- DOI: 10.1016/j.meegid.2015.06.023
Whole-genome sequencing of uropathogenic Escherichia coli reveals long evolutionary history of diversity and virulence
Abstract
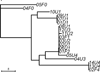
Uropathogenic Escherichia coli (UPEC) are phenotypically and genotypically very diverse. This diversity makes it challenging to understand the evolution of UPEC adaptations responsible for causing urinary tract infections (UTI). To gain insight into the relationship between evolutionary divergence and adaptive paths to uropathogenicity, we sequenced at deep coverage (190×) the genomes of 19 E. coli strains from urinary tract infection patients from the same geographic area. Our sample consisted of 14 UPEC isolates and 5 non-UTI-causing (commensal) rectal E. coli isolates. After identifying strain variants using de novo assembly-based methods, we clustered the strains based on pairwise sequence differences using a neighbor-joining algorithm. We examined evolutionary signals on the whole-genome phylogeny and contrasted these signals with those found on gene trees constructed based on specific uropathogenic virulence factors. The whole-genome phylogeny showed that the divergence between UPEC and commensal E. coli strains without known UPEC virulence factors happened over 32 million generations ago. Pairwise diversity between any two strains was also high, suggesting multiple genetic origins of uropathogenic strains in a small geographic region. Contrasting the whole-genome phylogeny with three gene trees constructed from common uropathogenic virulence factors, we detected no selective advantage of these virulence genes over other genomic regions. These results suggest that UPEC acquired uropathogenicity long time ago and used it opportunistically to cause extraintestinal infections.
Keywords: Next-generation sequencing; Phylogeny; Uropathogen.
Copyright © 2015 Elsevier B.V. All rights reserved.
Figures



Comment in
-
Re: Whole-Genome Sequencing of Uropathogenic Escherichia coli Reveals Long Evolutionary History of Diversity and Virulence.J Urol. 2017 Mar;197(3 Pt 1):706. doi: 10.1016/j.juro.2016.11.048. Epub 2016 Nov 11. J Urol. 2017. PMID: 28208529 No abstract available.
References
-
- Clark G, Paszkiewicz K, Hale J, Weston V, Constantinidou C, Penn C, Achtman M, McNally A. Genomic analysis uncovers a phenotypically diverse but genetically homogeneous Escherichia coli ST131 clone circulating in unrelated urinary tract infections. J. Antimicrob. Chemother. 2012;67:868–877. - PubMed
-
- Felsenstein J. PHYLIP (Phylogeny Inference Package) version 3.69. 2005
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

