Differential molecular response of monodehydroascorbate reductase and glutathione reductase by nitration and S-nitrosylation
- PMID: 26116026
- PMCID: PMC4566986
- DOI: 10.1093/jxb/erv306
Differential molecular response of monodehydroascorbate reductase and glutathione reductase by nitration and S-nitrosylation
Abstract
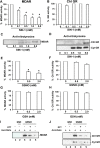
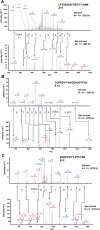
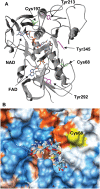
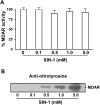
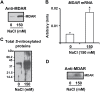
The ascorbate-glutathione cycle is a metabolic pathway that detoxifies hydrogen peroxide and involves enzymatic and non-enzymatic antioxidants. Proteomic studies have shown that some enzymes in this cycle such as ascorbate peroxidase (APX), monodehydroascorbate reductase (MDAR), and glutathione reductase (GR) are potential targets for post-translational modifications (PMTs) mediated by nitric oxide-derived molecules. Using purified recombinant pea peroxisomal MDAR and cytosolic and chloroplastic GR enzymes produced in Escherichia coli, the effects of peroxynitrite (ONOO(-)) and S-nitrosoglutathione (GSNO) which are known to mediate protein nitration and S-nitrosylation processes, respectively, were analysed. Although ONOO(-) and GSNO inhibit peroxisomal MDAR activity, chloroplastic and cytosolic GR were not affected by these molecules. Mass spectrometric analysis of the nitrated MDAR revealed that Tyr213, Try292, and Tyr345 were exclusively nitrated to 3-nitrotyrosine by ONOO(-). The location of these residues in the structure of pea peroxisomal MDAR reveals that Tyr345 is found at 3.3 Å of His313 which is involved in the NADP-binding site. Site-directed mutagenesis confirmed Tyr345 as the primary site of nitration responsible for the inhibition of MDAR activity by ONOO(-). These results provide new insights into the molecular regulation of MDAR which is deactivated by nitration and S-nitrosylation. However, GR was not affected by ONOO(-) or GSNO, suggesting the existence of a mechanism to conserve redox status by maintaining the level of reduced GSH. Under a nitro-oxidative stress induced by salinity (150mM NaCl), MDAR expression (mRNA, protein, and enzyme activity levels) was increased, probably to compensate the inhibitory effects of S-nitrosylation and nitration on the enzyme. The present data show the modulation of the antioxidative response of key enzymes in the ascorbate-glutathione cycle by nitric oxide (NO)-PTMs, thus indicating the close involvement of NO and reactive oxygen species metabolism in antioxidant defence against nitro-oxidative stress situations in plants.
Keywords: Glutathione reductase; S-nitrosoglutathione.; S-nitrosylation; monodehydroascorbate reductase; nitration; nitric oxide; peroxynitrite; reactive nitrogen species; salinity.
© The Author 2015. Published by Oxford University Press on behalf of the Society for Experimental Biology.
Figures

References
-
- Alvarez B, Radi R. 2003. Peroxynitrite reactivity with amino acids and proteins. Amino Acids 25, 295–311. - PubMed
-
- Asada K. 1992. Ascorbate peroxidase: a hydrogen peroxide-scavenging enzyme in plants. Physiologia Plantarum 85, 235–241.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

