Heparin interacts with elongation factor 1α of Cryptosporidium parvum and inhibits invasion
- PMID: 26129968
- PMCID: PMC4486996
- DOI: 10.1038/srep11599
Heparin interacts with elongation factor 1α of Cryptosporidium parvum and inhibits invasion
Abstract
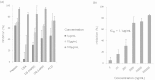
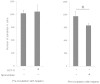
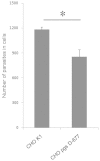
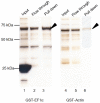
Cryptosporidium parvum is an apicomplexan parasite that can cause serious watery diarrhea, cryptosporidiosis, in human and other mammals. C. parvum invades gastrointestinal epithelial cells, which have abundant glycosaminoglycans on their cell surface. However, little is known about the interaction between C. parvum and glycosaminoglycans. In this study, we assessed the inhibitory effect of sulfated polysaccharides on C. parvum invasion of host cells and identified the parasite ligands that interact with sulfated polysaccharides. Among five sulfated polysaccharides tested, heparin had the highest, dose-dependent inhibitory effect on parasite invasion. Heparan sulfate-deficient cells were less susceptible to C. parvum infection. We further identified 31 parasite proteins that potentially interact with heparin. Of these, we confirmed that C. parvum elongation factor 1α (CpEF1α), which plays a role in C. parvum invasion, binds to heparin and to the surface of HCT-8 cells. Our results further our understanding of the molecular basis of C. parvum infection and will facilitate the development of anti-cryptosporidial agents.
Figures


Similar articles
-
Cryptosporidium parvum Elongation Factor 1α Participates in the Formation of Base Structure at the Infection Site During Invasion.J Infect Dis. 2020 May 11;221(11):1816-1825. doi: 10.1093/infdis/jiz684. J Infect Dis. 2020. PMID: 31872225 Free PMC article.
-
Elongation factor-1α is a novel protein associated with host cell invasion and a potential protective antigen of Cryptosporidium parvum.J Biol Chem. 2013 Nov 22;288(47):34111-34120. doi: 10.1074/jbc.M113.515544. Epub 2013 Sep 30. J Biol Chem. 2013. PMID: 24085304 Free PMC article.
-
Requirement of microtubules for secretion of a micronemal protein CpTSP4 in the invasive stage of the apicomplexan Cryptosporidium parvum.mBio. 2024 Feb 14;15(2):e0315823. doi: 10.1128/mbio.03158-23. Epub 2024 Jan 24. mBio. 2024. PMID: 38265238 Free PMC article.
-
Molecular targets for detection and immunotherapy in Cryptosporidium parvum.Biotechnol Adv. 2007 Jan-Feb;25(1):13-44. doi: 10.1016/j.biotechadv.2006.08.003. Epub 2006 Aug 30. Biotechnol Adv. 2007. PMID: 17055210 Review.
-
Host intestinal epithelial response to Cryptosporidium parvum.Adv Drug Deliv Rev. 2004 Apr 19;56(6):869-84. doi: 10.1016/j.addr.2003.10.034. Adv Drug Deliv Rev. 2004. PMID: 15063595 Review.
Cited by
-
Effect of Glycosaminoglycans on Cryptosporidium Oocyst Attachment and Excystation.Appl Environ Microbiol. 2023 Mar 29;89(3):e0173722. doi: 10.1128/aem.01737-22. Epub 2023 Feb 15. Appl Environ Microbiol. 2023. PMID: 36790186 Free PMC article.
-
Heparin Mimics Extracellular DNA in Binding to Cell Surface-Localized Proteins and Promoting Staphylococcus aureus Biofilm Formation.mSphere. 2017 Jun 21;2(3):e00135-17. doi: 10.1128/mSphere.00135-17. eCollection 2017 May-Jun. mSphere. 2017. PMID: 28656173 Free PMC article.
-
Antiparasitic Effects of Sulfated Polysaccharides from Marine Hydrobionts.Mar Drugs. 2021 Nov 12;19(11):637. doi: 10.3390/md19110637. Mar Drugs. 2021. PMID: 34822508 Free PMC article. Review.
-
Characterization of antigenic proteins of the Taenia solium postoncospheral form.Mol Biochem Parasitol. 2024 Sep;259:111621. doi: 10.1016/j.molbiopara.2024.111621. Epub 2024 May 3. Mol Biochem Parasitol. 2024. PMID: 38705360
-
A Single-Pass Type I Membrane Protein from the Apicomplexan Parasite Cryptosporidium parvum with Nanomolar Binding Affinity to Host Cell Surface.Microorganisms. 2021 May 8;9(5):1015. doi: 10.3390/microorganisms9051015. Microorganisms. 2021. PMID: 34066754 Free PMC article.
References
-
- Slapeta J. Cryptosporidiosis and Cryptosporidium Species in Animals and Humans: A Thirty Colour Rainbow? Int. J. Parasitol. 43, 957–970 (2013). - PubMed
-
- Thompson R. C. A. et al. Cryptosporidium and Cryptosporidiosis. Adv. Parasitol. 59, 77–158 (2005). - PubMed
-
- Odonoghue P. J. Cryptosporidium and Cryptosporidiosis in Man and Animals. Int. J. Parasitol. 25, 139–195 (1995). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

