ITS2 Secondary Structure Improves Discrimination between Medicinal "Mu Tong" Species when Using DNA Barcoding
- PMID: 26132382
- PMCID: PMC4488503
- DOI: 10.1371/journal.pone.0131185
ITS2 Secondary Structure Improves Discrimination between Medicinal "Mu Tong" Species when Using DNA Barcoding
Abstract
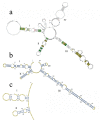
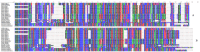
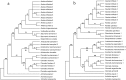
DNA barcoding is a promising species identification method, but it has proved difficult to find a standardized DNA marker in plant. Although the ITS/ITS2 RNA transcript has been proposed as the core barcode for seed plants, it has been criticized for being too conserved in some species to provide enough information or too variable in some species to align it within the different taxa ranks. We selected 30 individuals, representing 16 species and four families, to explore whether ITS2 can successfully resolve species in terms of secondary structure. Secondary structure was predicted using Mfold software and sequence-structure was aligned by MARNA. RNAstat software transformed the secondary structures into 28 symbol code data for maximum parsimony (MP) analysis. The results showed that the ITS2 structures in our samples had a common four-helix folding type with some shared motifs. This conserved structure facilitated the alignment of ambiguous sequences from divergent families. The structure alignment yielded a MP tree, in which most topological relationships were congruent with the tree constructed using nucleotide sequence data. When the data was combined, we obtained a well-resolved and highly supported phylogeny, in which individuals of a same species were clustered together into a monophyletic group. As a result, the different species that are often referred to as the herb "Mu tong" were successfully identified using short fragments of 250 bp ITS2 sequences, together with their secondary structure. Thus our analysis strengthens the potential of ITS2 as a promising DNA barcode because it incorporates valuable secondary structure information that will help improve discrimination between species.
Conflict of interest statement
Figures

References
-
- Baldwin BG, Sanderson MJ, Porter JM, Wojciechowski MF, Campbell CS, Donoghue MJ. The ITS region of nuclear ribosomal DNA: a valuable source of evidence on angiosperm phylogeny. Ann Mo Bot Gard.1995;82: 247–277.
-
- Alvarez I, Wendel JF. Ribosomal ITS sequences and plant phylogenetic inference. Mol Phylogenet Evol. 2003;29(3): 417–434. - PubMed
-
- Li DZ, Gao LM, Li HT, Wang H, Ge XJ, Liu JQ, et al. Comparative analysis of a large dataset indicates that internal transcribed spacer (ITS) should be incorporated into the core barcode for seed plants. Proc. Natl. Acad. Sci. USA 2011;108(49): 19641–19646. 10.1073/pnas.1104551108 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

